Carol Lefebvre, Julie Glanville, Simon Briscoe, Anne Littlewood, Chris Marshall, Maria-Inti Metzendorf, Anna Noel-Storr, Tamara Rader, Farhad Shokraneh, James Thomas and L. Susan Wieland on behalf of the Cochrane Information Retrieval Methods Group
This technical supplement should be cited as: Lefebvre C, Glanville J, Briscoe S, Littlewood A, Marshall C, Metzendorf M-I, Noel-Storr A, Rader T, Shokraneh F, Thomas J, Wieland LS. Technical Supplement to Chapter 4: Searching for and selecting studies. In: Higgins JPT, Thomas J, Chandler J, Cumpston MS, Li T, Page MJ, Welch VA (eds). Cochrane Handbook for Systematic Reviews of Interventions Version 6.1 (updated September 2020). Cochrane, 2020. Available from: www.training.cochrane.org/handbook.
This technical supplement is also available as a PDF document.
Throughout this technical supplement we refer to the Methodological Expectations of Cochrane Intervention Reviews (MECIR), which are methodological standards to which all Cochrane Protocols, Reviews, and Updates are expected to adhere. More information can be found on these standards at: https://methods.cochrane.org/mecir and, with respect to searching for and selecting studies, in Chapter 4 of the Cochrane Handbook for Systematic Review of Interventions.
1 Sources to search
For discussion of CENTRAL, MEDLINE and Embase as the key database sources to search, please refer to Chapter 4, Section 4.3. For discussion of sources other than CENTRAL, MEDLINE and Embase, please see the sections below.
1.1 Bibliographic databases other than CENTRAL, MEDLINE and Embase
1.1.1 The Cochrane Register of Studies
The Cochrane Register of Studies (CRS) is a bespoke Cochrane data repository and data management system, primarily used by Cochrane Information Specialists (CISs). The specialized trials registers maintained by CISs are stored and managed within the CRS. As such, it acts as a ‘meta-register’ of all the trials identified by Cochrane but each Cochrane Group has its own section (segment) within the larger database (Littlewood et al 2017). The Cochrane Central Register of Controlled Trials (CENTRAL) is created within the CRS, drawn partly from the references CISs add to their own segments and partly from references to trial reports sourced from other bibliographic databases (e.g. PubMed and Embase). The CRS is the only route available for publication of records in CENTRAL (Littlewood et al 2017).
As a piece of web-based software, the CRS provides tools to manage search activities both for the Cochrane group’s Specialized Register and for individual Cochrane Reviews. CISs are able to import records from external bibliographic databases and other sources into the CRS, de-duplicate them, share them with author teams and track what has been previously retrieved via searching and screened for each review. A further benefit is that trials register records (currently ClinicalTrials.gov and the WHO International Clinical Trials Registry Platform) are searchable from within the CRS. It is possible to store the full text of each bibliographic citation (and any accompanying documents, such as translations) within the CRS as an attachment but this should always be done in compliance with local copyright and database licensing agreements. Records added to the CRS that will be published in CENTRAL are automatically edited in accordance with the Cochrane HarmoniSR guidance, which ensures consistency in record formatting and output (HarmoniSR Working Group 2015).
The CRS captures links among references, studies and the Cochrane Reviews within which they appear. This information is drawn from CRS-D, a data repository which sits behind the CRS and includes all CENTRAL records, all included and excluded studies together with ongoing studies, studies awaiting classification and other records collected by CISs in their Specialized Registers. CRS-D has been designed to integrate with RevMan and Archie and this linking of data and information back to the reviews will ultimately help review teams find trials more efficiently. For example, CRS-D records can be linked to records in the Reviews Database that powers RevMan Web, so users can access additional data about the studies that appear in reviews, such as the characteristics of studies, ‘Risk of bias’ tables and, where possible, the extracted data from the study.
The CRS is a mixture of public records, i.e. CENTRAL records and private records for the use of Cochrane editorial staff only. Full access to the content in CRS is available only to designated staff within Cochrane editorial teams. Permission to perform tasks is controlled through Archie, Cochrane’s central server for managing documents and contact details (Littlewood et al 2017).
1.1.2 National and regional databases
In addition to MEDLINE and Embase, which are generally considered to be the key international general healthcare databases, many countries and regions produce bibliographic databases that focus on the literature produced in those regions and which often include journals and other literature not indexed elsewhere, such as African Index Medicus and LILACs (for Latin America and the Caribbean). It is highly desirable, for Cochrane reviews of interventions, that searches be conducted of appropriate national and regional bibliographic databases (MECIR C25). Searching these databases in some cases identifies unique studies that are not available through searching major international databases (Clark et al 1998, Brand-de Heer 2001, Clark and Castro 2001, Clark and Castro 2002, Abhijnhan et al 2007, Almerie et al 2007, Xia et al 2008, Atsawawaranunt et al 2009, Barnabas et al 2009, Manriquez 2009, Waffenschmidt et al 2010, Atsawawaranunt et al 2011, Wu et al 2013, Bonfill et al 2015, Cohen et al 2015, Xue et al 2016). Access to many of these databases is available free of charge. Others are only available by subscription or on a ‘pay-as-you-go’ basis. Indexing complexity and consistency varies, as does the sophistication of the search interfaces.
For a list of general healthcare databases, see Appendix.
1.1.3 Subject-specific databases
It is highly desirable, for authors of Cochrane reviews of interventions, to search appropriate subject specific bibliographic databases (MECIR C25). Which subject-specific databases to search in addition to CENTRAL, MEDLINE and Embase will be influenced by the topic of the review, access to specific databases and budget considerations.
Most of the main subject-specific databases such as AMED (alternative therapies), CINAHL (nursing and allied health) and PsycINFO (psychology and psychiatry) are available only on a subscription or ‘pay-as-you-go’ basis. Access to databases is, therefore, likely to be limited to those databases that are available to the Cochrane Information Specialist at the CRG editorial base or those that are available at the institutions of the review authors. Access arrangements vary according to institution. Review authors should seek advice from their medical / healthcare librarian or information specialist about access at their institution.
Although there is overlap in content coverage across Embase, MEDLINE and CENTRAL and subject-specific databases such as AMED, CINAHL and PsycINFO (Moseley et al 2009), their performance (Watson and Richardson 1999a, Watson and Richardson 1999b) and facilities vary. In addition, a comparison of British Nursing Index and CINAHL shows that even in databases in a specific field such as nursing, each database covers unique journal titles (Briscoe and Cooper 2014). To find qualitative research, CINAHL and PsycINFO should be searched in addition to MEDLINE and Embase (Subirana et al 2005, Wright et al 2015, Rogers et al 2017). Even in cases where research indicates low benefit in searching CINAHL, it is still suggested that for subject-specific reviews it should be considered as an option (Beckles et al 2013).
There are also several studies, each based on a single review, and therefore not necessarily generalizable to all reviews in all topics, showing that searching subject specific databases identified additional relevant publications. It is unclear, however, whether these additional publications would change the conclusions of the review. For example, for a review of exercise therapy for cancer patients, searching CancerLit, CINAHL, and PsycINFO identified additional records which were not retrieved by MEDLINE searches but searching SPORTDiscus identified no additional records (Stevinson and Lawlor 2004); for a review of social interventions, only four of the 69 (less than 6%) relevant studies were found by searching databases such as MEDLINE, while about half of the relevant studies were found by searching the Transport database (Ogilvie et al 2005); in an obesity review, searching the Health Management Information Consortium (HMIC) database identified about one fifth of included publications in addition to MEDLINE searches while CINAHL identified no new publications; and finally, in a tuberculosis review, searching CINAHL identified over 5% of the included publications in addition to MEDLINE, whereas the HMIC database identified no additional publications (Levay et al 2015).
For a list of subject-specific healthcare databases, see Appendix.
1.1.4 Citation indexes
Citation indexes are bibliographic databases which index citations in addition to the standard bibliographic content. They were originally developed to identify efficiently the reference lists of scholarly authors and the number of times a study or author is cited (Garfield 2007). Citation indexes can also be used creatively to identify studies which are similar to a source study, as it is probable that studies which cite or are cited by a source study will contain similar content.
Searching using a citation index is usually called ‘citation searching’ or ‘citation chasing’ and is further defined as ‘forwards citation searching’ or ‘backwards citation searching’ depending on which direction the citations are searched. Forwards citation searching identifies studies which cite a source study and backwards citation searching identifies studies cited by the source study. Citation indexes are mainly used for forwards citation searching, which is practically impossible to conduct manually, whereas backwards citation searching is relatively easy to conduct manually by consulting reference lists of source studies (see Section 1.3.4). Thus the focus in this section is on forwards citation searching. Citation indexes also facilitate author citation searching which is used to identify studies that are carried out by an author and studies that cite an author.
It is good practice to carry out forwards citation searching on studies that meet the eligibility criteria of a systematic review. Thus forwards citation searching usually takes place after the results of the bibliographic database searches have been screened and a set of potentially includable studies has been identified. Because citation searching is not based on pre-specified terminology it has the potential to retrieve studies that are not retrieved by the keyword-based search strategies that are conducted in bibliographic databases and other resources. This makes citation searching particularly effective in systematic reviews where the search terms are difficult to define, usefully extending to iterative citation searching of citations identified by citation searching (also known as ‘snowballing’) in some reported cases (Booth 2001, Greenhalgh and Peacock 2005, Papaioannou et al 2010, Linder et al 2015). Since researchers may selectively cite studies with positive results, forwards citation searching should be used with caution as an adjunct to other search methods in Cochrane Reviews.
There are varied findings on the efficiency of forwards citation searching, measured as the labour required to export and screen the results of searches relative to the number of unique studies identified (Wright et al 2014, Hinde and Spackman 2015, Levay et al 2016, Cooper et al 2017a). Most studies, however, which compared the results of forwards citation searching with other search methods found that citation searching identified one or more unique studies which were relevant to the review question (Greenhalgh and Peacock 2005, Papaioannou et al 2010, Wright et al 2014, Hinde and Spackman 2015, Linder et al 2015). Reviews of recently published studies, such as review updates, are less likely to benefit from forwards citation searching than reviews with no historical date limit for includable studies due to the relatively limited time for recent studies to be cited. When conducting a review update, however, searchers should consider carrying out forwards citation searching on the studies included in the original review and on the original review itself.
The two main subscription citation indexes are Web of Science, which was launched in 1964 and is currently provided by Clarivate Analytics, and Scopus, which was launched in 2004 by Elsevier. Google Scholar, which was also launched in 2004, can be used for forwards but not backwards citation searching. Microsoft Academic was relaunched in 2015 (Sinha et al 2015). It can be used for both forwards and backwards citation searching. A summary of each resource is provided below. There are published comparative studies which can be consulted for a more detailed analysis (Kulkarni et al 2009, Wright et al 2014, Levay et al 2016, Cooper et al 2017b).
Web of Science
Web of Science (formerly Web of Knowledge), produced by Clarivate Analytics, comprises several databases. The ‘Core Collection’ databases cover the sciences (1900 to date), social sciences (1956 to date), and arts and humanities (1975 to date). The sciences and social sciences collections are divided into journal articles and conference proceedings, which can be searched separately. In total, the Web of Science Core Collection contains over 74 million records from more than 21,100 journal titles, books and conference proceedings (Web of Science 2019). Additional databases are available via the Web of Science platform, also on a subscription basis. Author citation searching is possible in Web of Science but it does not automatically distinguish between authors with the same name unless they have registered for a uniquely assigned Web of Science ResearcherID.
https://clarivate.com/products/web-of-science/
Scopus
Scopus, produced by Elsevier, covers health sciences, life sciences, physical sciences and social sciences. As of March 2019, it contains approximately 69 million records from 21,500 journal titles and 88,800 conferences proceedings dating back to 1823 (Scopus 2017). Citation details are mainly available from 1996 to date, though Scopus is in the process of adding details of pre-1996 citations and is expanding the total number of pre-1996 records (Beatty 2015). A unique identification number is automatically assigned to each author in the database which enables it to distinguish between authors with the same names when author citation searching. Errors are still possible, however, as publications are not always assigned correctly to author ID numbers and authors are sometimes erroneously assigned more than one ID number.
https://www.elsevier.com/solutions/scopus
Google Scholar
Google Scholar is a freely available scholarly search engine which uses automated web crawlers to identify and index scholarly references, including published studies and grey literature. Although it can only be used for forwards citation searching, this limitation has little practical significance as backwards citation searching can be easily conducted manually by checking reference lists. The precise number of journals indexed by Google Scholar is not known because it does not use a pre-specified list of journals to populate its content. There is, however, evidence that it has sufficient citation coverage to be used as an alternative to Web of Science or Scopus, if these databases are not available (Wright et al 2014, Levay et al 2016).
A disadvantage of Google Scholar’s automated study identification method is that it produces more duplicate citations than Web of Science, which indexes pre-specified journal content (Haddaway et al 2015). Scopus, which uses a similar indexing method to Web of Science, is also likely to produce fewer duplicates than Google Scholar. A further disadvantage of Google Scholar is that the export features are basic and inefficient and are only marginally improved by linking to its preferred reference manager software, Zotero (Bramer et al 2013, Levay et al 2016). Google Scholar citations can also be exported to the Publish or Perish software (Harzing 2007). Finally, Google Scholar limits the number of viewable results to 1000 and does not disclose how the top 1000 results are selected, thus compromising the transparency and reproducibility of search results (Levay et al 2016).
Microsoft Academic
Microsoft Academic is a relatively new scholarly search engine, with many similarities to Google Scholar. It is free to access, and identifies its source material from the ‘Bing’ web crawler, and so contains both journal articles and reports of research that are not indexed in mainstream bibliographic databases. Like Google Scholar, it is made up of a ‘graph’ of publications that are connected to one another by citation, author, and institutional relationships. Unlike Google Scholar, it provides for both forwards and backwards citation searching, and also contains a ‘related’ documents feature, which identifies documents which its algorithm considers to be closely related to one another. As well as being available through its website, Microsoft Academic also publishes an Application Programming Interface (API) - for other software applications to ‘plug’ into - and it is possible to obtain copies of the entire dataset on request. The API and raw data are probably of greater interest to tool developers than information specialists (though there are some tools in R that provide access to the API), but the greater openness of this dataset compared with Google Scholar may result in the development of a number of useful applications for systematic review authors over time.
https://academic.microsoft.com/
Web of Science, Scopus, Google Scholar and Microsoft Academic all provide wide coverage of healthcare journal publications. There are, however, differences in the number of records indexed in each citation index and in the methods used to index records, and there is evidence that these differences affect the number of citations which are identified when citation searching (Kulkarni et al 2009, Wright et al 2014, Rogers et al 2016). It is not a requirement for Cochrane Reviews, however, to conduct exhaustive citation searching using multiple citation indexes. Review authors and information specialists should consider the time and resources available and the likelihood of identifying unique studies for the review question, when planning whether and how to conduct forwards citation searching.
Further evidence-based analysis of the value of citation searching for systematic reviews can be found on the regularly updated SuRe Info portal in the section entitled Value of using different search approaches (http://vortal.htai.org/?q=node/993).
1.1.5 Dissertations and theses databases
It is highly desirable, for authors of Cochrane reviews of interventions, to search relevant grey literature sources such as reports, dissertations, theses, databases and databases of conference abstracts (MECIR C28). Dissertations and theses are a subcategory of grey literature, which may report studies of relevance to review authors. Searching for unpublished academic research may be important for countering possible publication bias but it can be time consuming and in some cases yield few included studies (van Driel et al 2009). In some areas of medicine, searching for and retrieving unpublished dissertations has been shown to have a limited influence on the conclusions of a review (Vickers and Smith 2000, Royle et al 2005). In other areas of medicine, however, it is essential to broaden the search to include unpublished trials, for example in oncology and in complementary medicine (Egger et al 2003). In a study of 129 systematic reviews from three Cochrane Review Groups (the Acute Respiratory Infections Group, the Infectious Diseases Group and the Developmental, Psychosocial and Learning Problems Group) there was wide variation in the retrieval and inclusion of dissertations (Hartling et al 2017). It is possible that a study which would affect the conclusions would be missed if the search is not comprehensive enough to include searches for unpublished trials including those reported only in dissertation and theses (Egger et al 2003). The failure to search for unpublished trials, such as those in dissertation and theses databases may lead to biased results in some reviews (Ziai et al 2017). Dissertations and theses are not normally indexed in general bibliographic databases such as MEDLINE or Embase, but there are exceptions, such as CINAHL, which indexes nursing, physical therapy and occupational health dissertations and PsycINFO, which indexes dissertations in psychiatry and psychology.
To identify relevant studies published in dissertations or theses it is advisable to search specific dissertation sources:
- The US-based Center for Research Libraries (CRL) is an international consortium of university, college, and independent research libraries (http://catalog.crl.edu/search~S4)
- The LILACS database includes some theses and dissertations from Latin American and Caribbean countries (http://lilacs.bvsalud.org/en/)
- Open Access Theses and Dissertations (OATD) includes electronic theses and databases that are free to access and read online from participating universities from around the world (https://oatd.org/)
- ProQuest Dissertations and Theses Global (PQDT) is the best-known commercial database for searching dissertations. Access to PQDT is by subscription. As at August 2019, ProQuest Dissertations and Theses Global database indexes approximately 5 million doctoral dissertations and Master’s theses from around the world (http://www.proquest.com/products-services/pqdtglobal.html)
Other sources of dissertations and theses include the catalogues and resources produced by national libraries and research centres, for example:
- Australian theses are searchable via the National Library of Australia’s Trove service (http://trove.nla.gov.au/)
- DART-Europe is a partnership of several research libraries and library consortia which provides global access to European research theses via a portal. A list of institutions, national libraries and consortia who contribute to the portal can be found here: (http://www.dart-europe.eu/basic-search.php)
- Deutsche Nationalbibliothek (German National Library) provides access to electronic versions of theses and dissertations since 1998 (https://www.dnb.de/dissonline)
-
The Networked Digital Library of Theses and Dissertations (NDLTD) is an international organization dedicated to promoting the adoption, creation, use, dissemination, and preservation of electronic theses and dissertations.
(http://search.ndltd.org/) - Swedish University Dissertations offers dissertations in English, some of which are available to download (http://www.dissertations.se/)
- Theses Canada provides access to the National Library of Canada’s records of PhD and Master’s theses from Canadian universities (www.collectionscanada.gc.ca/thesescanada/)
Other countries also offer access to dissertations and theses in their national languages.
Whenever possible, review authors should attempt to include all relevant studies of acceptable quality, irrespective of the type of publication, since the inclusion of these may have an impact in situations where there are few relevant studies, or where there may be vested interests in the published literature (Hartling et al 2017). The inclusion of unpublished trials will increase precision, generalizability and applicability of findings (Egger et al 2003). In the interest of feasibility, review authors should assess their research questions and topic area, and seek advice from content experts when selecting dissertation and theses databases to search. Review authors should consult their Cochrane Information Specialist, local library or university for information about dissertations and theses databases in their country or region.
1.1.6 Grey literature databases
As stated above, it is highly desirable, for authors of Cochrane reviews of interventions, to search relevant grey literature sources such as reports, dissertations, theses, databases and databases of conference abstracts (MECIR C28).
Grey literature was defined at GL3, the Third International Conference on Grey Literature on 13 November 1997 in Luxembourg as “that which is produced on all levels of government, academics, business and industry in print and electronic formats, but which is not controlled by commercial publishers” (Farace and Frantzen 1997). On 6 December 2004, at GL6, the Sixth Conference in New York City, a clarification was added: grey literature is “... not controlled by commercial publishers, i.e. where publishing is not the primary activity of the producing body …” (Farace and Frantzen 2005). In a 2017 audit of 203 systematic reviews published in high-impact general medical journals in 2013, 64% described an attempt to search for unpublished studies. The audit showed that reviews published in the Cochrane Database of Systematic Reviews were significantly more likely to include a search for grey literature than those published in standard journals (Ziai et al 2017). A Cochrane Methodology Review indicated that published trials showed an overall greater treatment effect than grey literature trials (Hopewell et al 2007a). Although failure to identify trials reported in conference proceedings and other grey literature might affect the results of a systematic review (Hopewell et al 2007a), a recent systematic review showed that this was only the case in a minority of reviews (Schmucker et al 2017). Since the impact of excluding unpublished data is unclear, review authors should consider the time and effort spent when planning the grey-literature portion of the search.
Grey literature’s diverse formats and audiences can present a significant challenge in a systematic search for evidence. Locating grey literature can often be challenging, requiring librarians to use several databases from various host providers or websites, some of which they may not be familiar with (Saleh et al 2014, Haddaway and Bayliss 2015). There are many characteristics of grey literature that make it difficult to search systematically. Further, there is no ‘gold standard’ for rigorous systematic grey literature search methods and few resources on how to conduct this type of search (Godin et al 2015, Paez 2017). One challenge of searching the grey literature is managing an abundance of material. Often, there are many sources to search but some authors of very broad or cross-disciplinary topics may find it necessary to impose some limits on the extent of their grey literature searching by considering what is feasible within limited time and resources (Mahood et al 2014). For example, since nearly half of the citations found in reviews of new and emerging non-drug technologies are grey literature, searchers should consider focusing their efforts on search engines and aggregator sites to increase feasibility (Farrah and Mierzwinski-Urban 2019). Google Scholar can help locate a large volume of grey literature and specific, known studies, however, it should not be used as the only resource for systematic review searches (Haddaway et al 2015). The types of grey literature that are useful in specific reviews may depend on the research question and researchers may decide to tailor the search to the question (Levay et al 2015). For example, unpublished academic research may be important for countering possible publication bias and can be targeted via specific repositories for preprints, theses and funding registries. Alternatively, if the research question is related to implementation or if the researchers are interested in material to support their implications for practice section, then organizational reports, government documents and monitoring and evaluation reports, might be important for ensuring the search is extensive and fit for purpose (Haddaway and Bayliss 2015).
Careful documentation throughout the search process will demonstrate that efforts have been made to be comprehensive and will help in making the grey literature searching as reproducible as possible (Stansfield et al 2016).
The following resources can help authors plan a manageable and thorough approach to searching the grey literature for their topic.
- The Canadian Agency for Drugs and Technologies in Health (CADTH) publishes a resource entitled ‘Grey Matters: a practical tool for searching health-related grey literature’ (https://www.cadth.ca/resources/finding-evidence/grey-matters) which lists a considerable number of grey literature sources together with annotations about their content as well as search hints and tips.
- GreySource (http://greynet.org/greysourceindex.html) provides links to self-described sources of grey literature. Only web-based resources that explicitly refer to the term grey literature (or its equivalent in any language) are listed. The links are categorized by subject, so that authors can quickly identify relevant sources to pursue.
- The Health Management Information Consortium (HMIC) Database (https://www.kingsfund.org.uk/consultancy-support/library-services) contains records from the Library and Information Services department of the UK Department of Health and the King’s Fund Information and Library Service. It includes all UK Department of Health publications including circulars and press releases. The King’s Fund is an independent health charity that works to develop and improve management of health and social care services. The database is considered to be a good source of grey literature on topics such as health and community care management, organizational development, inequalities in health, user involvement, and race and health.
- The US National Technical Information Service (NTIS; www.ntis.gov) provides access to the results of both US and non-US government-sponsored research and can provide the full text of the technical report for most of the results retrieved. NTIS is free of charge on the internet and goes back to 1964.
- OpenGrey (www.opengrey.eu) is a multidisciplinary European grey literature database, covering science, technology, biomedical science, economics, social science and humanities. Each record has an English title and / or English keywords. Some records include an English abstract (starting in 1997). The database includes technical or research reports, doctoral dissertations, conference presentations, official publications, and other types of grey literature. Information is also provided regarding how to access the documents included in the database.
- PsycEXTRA (http://www.apa.org/pubs/databases/psycextra/) is a companion database to PsycINFO in psychology, behavioural science and health. It includes references from newsletters, magazines, newspapers, technical and annual reports, government reports and consumer brochures. PsycEXTRA is different from PsycINFO (https://www.apa.org/pubs/databases/psycinfo/index) in its format, because it includes abstracts and citations plus full text for a major portion of the records. There is no coverage overlap between PsycEXTRA and PsycINFO.
Conference abstracts are a particularly important source of grey literature and are further covered in Section 1.3.3.
1.2 Ongoing studies and unpublished data sources: further considerations
This section should be read in conjunction with Chapter 4, Sections 4.3.2, 4.3.3, and 4.3.4.
1.2.1 Trials registers and trials results registers
It is mandatory, for authors of Cochrane reviews of interventions, to search trials registers and repositories of results, where relevant to the topic, through ClinicalTrials.gov, the WHO International Clinical Trials Registry Platform (ICTRP) portal and other sources as appropriate (MECIR C27) (see Chapter 4, Section 4.3.3). Although ClinicalTrials.gov is included as one of the registers within the WHO ICTRP portal, it is recommended that both ClinicalTrials.gov and the ICTRP portal are searched separately, from within their own interfaces, due to additional features in ClinicalTrials.gov (Glanville et al 2014)(see below).
Several initiatives have led to the development of and recommendations to search trials registers. The International Committee of Medical Journal Editors (ICMJE) requires prospective registration of studies for subsequent publication in their journals, and there is a legal requirement that the results of certain studies must be posted within a given timeframe. Several studies have shown, however, that adherence to these requirements is mixed (Gill 2012, Huser and Cimino 2013b, Huser and Cimino 2013a, Jones et al 2013, Anderson et al 2015, Dal-Re et al 2016, Goldacre et al 2018, Jorgensen et al 2018) and that results posted on ClinicalTrials.gov show discordance when compared with results published in journal articles (Gandhi et al 2011, Earley et al 2013, Hannink et al 2013, Becker et al 2014, Hartung et al 2014, De Oliveira et al 2015) or both of the above (Jones and Platts-Mills 2012, Adam et al 2018).
ClinicalTrials.gov
In February 2000, the US National Library of Medicine (NLM) launched ClinicalTrials.gov (https://clinicaltrials.gov/ct2/home). ClinicalTrials.gov was created as a result of the Food and Drug Administration Modernization Act of 1997 (FDAMA). FDAMA required the U.S. Department of Health and Human Services, through the US National Institutes of Health (NIH), to establish a registry of clinical trials information for both (US) federally and privately funded trials conducted under ‘investigational new drug’ applications to test the effectiveness of experimental drugs for “serious or life-threatening diseases or conditions”. The ClinicalTrials.gov registration requirements were expanded after the US Congress passed the FDA Amendments Act of 2007 (FDAAA). Section 801 of FDAAA (FDAAA 801) required more types of trials to be registered and additional trial registration information to be submitted. The law also required the submission of results for certain trials. This led to the expansion of ClinicalTrials.gov to include information on study participants and a summary of study outcomes, including adverse events. Results have been made available since September 2008. Further legislation has expanded the coverage of results in ClinicalTrials.gov, which now serves as a major international register including clinical trials conducted across over 200 countries. Searches of ClinicalTrials.gov can be limited to studies which include results by selecting ‘Studies With Results’ from the pull-down menu at the ‘Study Results’ option on the Advanced Search page (https://clinicaltrials.gov/ct2/search/advanced). Research has shown that the most reliable way of searching ClinicalTrials.gov is to conduct a highly sensitive ‘single concept’ search in the basic interface of ClinicalTrials.gov (Glanville et al 2014). This study also suggested that use of the advanced interface seemed to improve precision without loss of sensitivity and this interface might be preferred when large numbers of search results are anticipated.
Search help for ClinicalTrials.gov is available from the following links:
How to Use Basic Search
https://clinicaltrials.gov/ct2/help/how-find/basic
How to Use Advanced Search
https://clinicaltrials.gov/ct2/help/how-find/advanced
How to Read a Study Record
https://clinicaltrials.gov/ct2/help/how-read-study
How to Use Search Results
https://clinicaltrials.gov/ct2/help/how-use-search-results
The World Health Organization International Clinical Trials Registry Platform search portal (WHO ICTRP)
In May 2007, the World Health Organization (WHO) launched the International Clinical Trials Registry Platform (ICTRP) search portal (http://apps.who.int/trialsearch/), to search across a range of trials registers, similar to the initiative launched some years earlier by Current Controlled Trials with their ‘metaRegister’ (which has ceased publication). Currently (August 2019), the WHO portal searches across 17 registers (including ClinicalTrials.gov but note the guidance above regarding searching ClinicalTrials.gov separately through the ClinicalTrials.gov interface). Research has shown that the most reliable way of searching the ICTRP is to conduct a highly sensitive ‘single concept’ search in the ICTRP basic interface (Glanville et al 2014). This study suggested that use of the ICTRP advanced interface might be problematic because of reductions in sensitivity.
Search help for the ICTRP is available from the following link:
http://apps.who.int/trialsearch/tips.aspx
Other trials registers
HSRProj (Health Services Research Projects in Progress) (https://hsrproject.nlm.nih.gov/) provides information about ongoing health services research and public health projects. It contains descriptions of research in progress funded by US federal and private grants and contracts for use by policy makers, managers, clinicians and other decision makers. It provides access to information about health services research in progress before results are available in a published form.
Many countries and regions maintain trials results registers. There are also many condition-specific trials registers, especially in the field of cancer, which are too numerous to list. Some pharmaceutical companies make available information about their clinical trials through their own websites, either instead of or in addition to the information they make available through national or international websites.
In addition, Clinical Trial Results (www.clinicaltrialresults.org) is a website that hosts slide and video presentations from clinical trialists, especially in the field of cardiology but also other specialties, reporting the results of clinical trials.
Further listings of international, national, regional, subject-specific and industry trials registers, together with guidance on how to search them can be found on a website developed in 2009 by two of the co-authors of this chapter (JG and CL) entitled Finding clinical trials, research registers and research results (https://sites.google.com/a/york.ac.uk/yhectrialsregisters/).
1.2.2 Regulatory agency sources and clinical study reports
The EU Clinical Trials Register (EUCTR)
The EUCTR contains protocol and results information for interventional clinical trials on medicines conducted in the European Union (EU) and the European Economic Area (EEA) which started after 1 May 2004. It enables searching for information in the EudraCT database, which is used by national medicines regulators for data related to clinical trial protocols. Results data are extracted from data entered by the sponsors into EudraCT. The EUCTR has been a ‘primary registry’ in the ICTRP since September 2011 but in the absence of any evidence to the contrary, it is recommended that searches of the EUCTR should be carried out within the EUCTR and not solely within the ICTRP (in line with the advice above regarding searching ClinicalTrials.gov). The register currently (August 2019) contains information about approximately 60,000 clinical trials. Searches can be limited to ‘Trials with results’ under the ‘Results status’ option and up to 50 records can be downloaded at a time.
https://www.clinicaltrialsregister.eu/ctr-search/search
Drugs@FDA, OpenTrialsFDA Prototype and medical devices
Drugs@FDA is hosted by the US Food and Drug Administration and provides information about most of the drugs approved in the US since 1939. For those approved more recently (from 1998), there is often a ‘Review’, which contains the scientific analyses that provided the basis for approval of the new drug. In 2012, new search options were introduced, enabling search strategies to be saved and re-run and results to be downloaded to a spreadsheet (Goldacre et al 2017).
(http://www.accessdata.fda.gov/scripts/cder/daf/)
The OpenTrialsFDA Prototype initiative makes data from FDA documents (Drug Approval Packages) more easily accessible and searchable, links the data to other clinical trial data and presents the data through a new user-friendly web interface
(https://opentrials.net/opentrialsfda/)
The FDA also makes information about devices, including several medical device databases (including the Post-Approval Studies (PAS) Database and a database of Premarket Approvals (PMA)), available on its website at:
Clinical study reports
Clinical study reports (CSRs) are reports of clinical trials, which provide detailed information on the methods and results of clinical trials submitted in support of marketing authorization applications. Cochrane recently funded a project under the Methods Innovation Funding programme to draft interim guidance to help Cochrane review authors decide whether to include data from clinical study reports (CSRs) and other regulatory documents in a Cochrane Review.
http://methods.cochrane.org/methods-innovation-fund-2. (Hodkinson et al 2018, Jefferson et al 2018)
A Clinical Study Reports Working Group has been established in Cochrane to take this work forward and to consider how CSRs might be used in Cochrane Reviews in future. To date, only one Cochrane Review is based solely on CSRs, that is the 2014 review update on neuraminidase inhibitors for preventing and treating influenza in healthy adults and children (Jefferson et al 2014).
In late 2010, the European Medicines Agency (EMA) began releasing CSRs (on request) under their Policy 0043. In October 2016, they began to release CSRs under their Policy 0070. The policy applies only to documents received since 1 January 2015. CSRs are available for approximately 150 products (as at September 2019) (https://clinicaldata.ema.europa.eu/web/cdp/background).
In order to download the full CSR documents, it is necessary to register for use “for academic and other non-commercial research purposes” and to provide an email address and a place of address in the European Union, or provide details of a third party, resident or domiciled in the European Union, who will be considered to be the user.
https://clinicaldata.ema.europa.eu/web/cdp/termsofuse
The FDA does not currently routinely provide access to CSRs, only their own internal reviews, as noted above. In January 2018, however, they announced a voluntary pilot programme to disclose up to nine recently approved drug applications, limited to CSRs for the key ‘pivotal’ trials that underpin drug approval (Doshi 2018). A public consultation of this pilot project (which included only one CSR) was undertaken in August 2019.
The Japanese Pharmaceuticals and Medical Devices Agency (PMDA) also provides access to its own internal reviews of approved drugs and medical devices but not the original CSRs. These can be found in the Reviews section of its website at:
https://www.pmda.go.jp/english/review-services/reviews/0001.html
https://www.pmda.go.jp/english/review-services/reviews/approved-information/drugs/0001.html
In April 2019 Health Canada announced that it was starting to make clinical information about drugs and devices publicly available on its website (https://clinical-information.canada.ca/search/ci-rc) (Lexchin et al 2019). As at August 2019, information was available for 10 drug records and three medical device records.
1.3 Journals and other non-bibliographic database sources
1.3.1 Handsearching
Handsearching involves a manual page-by-page examination of the entire contents of a journal issue or conference proceedings to identify all eligible reports of trials. (For discussion of ‘handsearching’ full-text journals available electronically, see Section 1.3.2) In journals, reports of trials may appear in articles, abstracts, news columns, editorials, letters or other text. Handsearching healthcare journals and conference proceedings can be a useful adjunct to searching electronic databases for at least two reasons: 1) not all trial reports are included in electronic bibliographic databases, and 2) even when they are included, they may not contain relevant search terms in the titles or abstracts or be indexed with terms that allow them to be easily identified as trials (Dickersin et al 1994). It should be noted, however, that handsearching is not a requirement for all Cochrane Reviews and review authors should seek advice from their Cochrane Information Specialist or their medical / healthcare librarian or information specialist with respect to whether handsearching might be valuable for their review, and if so, what to search and how (Littlewood et al 2017). Each journal year or conference proceeding that is to be handsearched should be searched thoroughly and competently by a well-trained handsearcher, ideally for all reports of trials, irrespective of topic, so that once it has been handsearched it will not need to be searched again. A Cochrane Methodology Review found that a combination of handsearching and electronic searching is necessary for full identification of relevant reports published in journals, even for those that are indexed in MEDLINE (Hopewell et al 2007b). This was especially the case for articles published before 1991 when there was no indexing term for randomized trials in MEDLINE and for those articles that are in parts of journals (such as supplements and conference abstracts) which are not routinely indexed in databases such as MEDLINE. Richards’ review (Richards 2008) found that handsearching was valuable for finding trials reported in abstracts or letters, or in languages other than English. We note that Embase is now a good source of conference abstracts.
To facilitate the identification of all published trials, Cochrane has organized extensive handsearching efforts. Over 3000 journals have been, or are being, searched within Cochrane. The list of journals that have already been handsearched, with the dates of the search and whether the search has been completed is available via the Handsearched Journals tab in the Cochrane Register of Studies Online at crso.cochrane.org, (Cochrane Account login required). Cochrane Information Specialists can edit records of journals that are being handsearched and can add new handsearch records to the Register (Littlewood et al 2017). Since many conference proceedings are now included within Embase, the information specialist will also check coverage of specific conferences of interest by checking the Embase list of conferences (https://www.elsevier.com/solutions/embase-biomedical-research/embase-coverage-and-content). Handsearching should still be considered, however, since searches of Embase will not necessarily find all the trials records in a conference issue (Stovold and Hansen 2011).
Cochrane groups and authors can prioritize handsearching based on where they expect to identify the most trial reports. This prioritization can be informed by searching CENTRAL, MEDLINE and Embase in a topic area and identifying which journals appear to be associated with the most retrieved citations. Preliminary evidence suggests that most of the journals with a high yield of trial reports are indexed in MEDLINE (Dickersin et al 2002) but this may reflect the fact that Cochrane contributors have concentrated early efforts on searching these journals. Therefore, journals not indexed in MEDLINE or Embase should also be considered for handsearching. Research into handsearching journals in a range of languages suggests that handsearching journals published in languages other than English is still helpful for identifying trials which have not been retrieved by database searches (Blumle and Antes 2005, Fedorowicz et al 2005, Al-Hajeri et al 2006, Nasser and Al Hajeri 2006, Chibuzor and Meremikwu 2009). The value of handsearching may vary from topic to topic. In physical therapy and respiratory disease, recent studies have found handsearching yielded additional studies (Stovold and Hansen 2011, Craane et al 2012). Identifying studies of handsearching in specific disease areas may help to inform decisions around handsearching.
The Cochrane Training Manual for Handsearchers is available on the Cochrane Information Retrieval Methods Group Website: http://methods.cochrane.org/irmg/resources.
1.3.2 Full text journals available electronically
The full text of many journals is available electronically on the internet. Access may be partially or wholly on a subscription basis or free of charge. In addition to providing a convenient method for retrieving the full article of already identified records, full-text journals can also be searched electronically, depending on the search interface, by entering relevant keywords in a similar way to searching for records in a bibliographic database. Electronic journals can also be ‘handsearched’ in a similar manner to that advocated for journals in print form, in that each screen or ‘page’ can be checked for possibly relevant studies in the same way as handsearching a print journal (see Section 1.3.1). When reporting handsearching, it is important to specify whether the full text of a journal has been searched electronically or using the print version. Some journals omit sections of the print version, for example letters, from the electronic version and some include supplementary information such as extra articles in the electronic format only.
Most academic institutions subscribe to a wide range of electronic journals and these are therefore available free of charge at the point of use to members of those institutions. Review authors should seek advice about electronic journal access from the library service at their institution. Some professional organizations provide access to a range of journals as part of their membership package. In some countries similar arrangements exist for health service employees through national licences.
Several international initiatives provide free or low-cost online access to full-text journals (and databases). The Health InterNetwork Access to Research Initiative (HINARI) provides access to approximately 15,000 journals (and up to 60,000 e-books), in 30 different languages, to health institutions in more than 120 low and middle income countries, areas and territories (World Health Organization 2019). Other initiatives include the International Network for the Availability of Scientific Publications (INASP) and Electronic Information for Libraries (EIFL).
A local electronic or print copy of any possibly relevant article found electronically in a subscription journal should be taken and filed (within copyright legislation), as the subscription to that journal may cease. The same applies to electronic journals available free of charge, as the circumstances around availability of specific journals might change. We have not been able to identify any research evidence regarding searching full-text journals available electronically. Authors are not routinely expected to search full-text journals available electronically for their reviews, but they should discuss with their Cochrane Information Specialist whether, in their particular case, this might be beneficial.
1.3.3 Conference abstracts and proceedings
It is highly desirable, for authors of all Cochrane reviews of interventions, to search relevant databases of conference abstracts (MECIR C28). Although conference proceedings are not indexed in MEDLINE, about 2.5 million conference abstracts from about 7,000 conferences (as at August 2019) are now indexed in Embase.
Elsevier provides a list of conferences it indexes in Embase, as mentioned above: (https://www.elsevier.com/solutions/embase-biomedical-research/embase-coverage-and-content). As a result of Cochrane’s Embase project (see Section 2.1.2), conference abstracts that are indexed in Embase and are reports of RCTs are now being included in CENTRAL. Other resources such as the Web of Science Conference Proceedings Citation Indexes also include conference abstracts. A Cochrane Methodology Review found that trials with positive results tended to be published in approximately 4 to 5 years whereas trials with null or negative results were published after about 6 to 8 years (Hopewell et al 2007c) and not all conference presentations are published or indexed (Slobogean et al 2009). Over one-half of trials reported in conference abstracts never reach full publication (Diezel et al 1999, Scherer et al 2018) and those that are eventually published in full have been shown to have results that are systematically different from those that are never published in full (Scherer et al 2018). In addition, conference abstracts / proceedings are a good source to track disagreements between the original abstract and the full report of studies (Chokkalingam et al 1998, Pitkin et al 1999). Additionally, trials with positive findings are more likely to be published than those which do not have positive findings (Salami and Alkayed 2013). It is, therefore, important to try to identify possibly relevant studies reported in conference abstracts through specialist database sources and by searching those abstracts that are made available on the internet, on CD-ROM / DVD or in print form. Many conference proceedings are published as journal supplements or as proceedings on the website of the conference or the affiliated organization.
1.3.4 Other reviews, guidelines and reference lists as sources of studies
It is highly desirable, for authors of Cochrane reviews of interventions, to search within previous reviews on the same topic (MECIR C29) and it is mandatory, for authors of Cochrane reviews of interventions, to check reference lists of included studies and any relevant systematic reviews identified (MECIR C30). Reviews can provide relevant studies and references, and may also provide information about the search strategy used, which may inform the current review (Hunt and McKibbon 1997, Glanville and Lefebvre 2000). Copies of previously published reviews on, or relevant to, the topic of interest should be obtained and checked for references to the included (and excluded) studies. Various sources for identifying previously published reviews are described below.
As well as the Cochrane Database of Systematic Reviews (CDSR), until recently, the Cochrane Library included the Database of Abstracts of Reviews of Effects (DARE) and the Health Technology Assessment Database (HTA Database), produced by the Centre for Reviews and Dissemination (CRD) at the University of York in the UK. Both databases provide information on published reviews of the effects of health care (Petticrew et al 1999). Searches of MEDLINE, Embase, CINAHL, PsycINFO and PubMed to identify candidate records were continued until the end of 2014 and bibliographic records were published on DARE until 31 March 2015. CRD will maintain secure archive versions of DARE until at least 2021. CRD continued to maintain and add records to the HTA database until 31 March 2018. It is being taken over by The International Network of Agencies for Health Technology Assessment (INAHTA) https://www.crd.york.ac.uk/CRDWeb/. Since 1 April 2015 the NIHR Dissemination Centre at the University of Southampton has had summaries of new research available. Details can be found at http://www.disseminationcentre.nihr.ac.uk/.
KSR Evidence, a subscription database, aims to include all systematic reviews and meta-analyses published since 2015 (https://ksrevidence.com/). KSR Evidence was developed by Kleijnen Systematic Reviews Ltd (KSR) (www.systematic-reviews.com). KSR produces and disseminates systematic reviews, cost-effectiveness analyses and health technology assessments of research evidence in health care. The database also includes an advanced search option, suitable for information specialists.
CRD provides an international register of prospectively registered systematic reviews in health and social care called PROSPERO, which (as at August 2019) contained over 50,000 records (www.crd.york.ac.uk/prospero/) (Page et al 2018). Key features from the review protocol are recorded and maintained as a permanent record. PROSPERO aims to provide a comprehensive listing of systematic reviews registered at inception to help avoid duplication and reduce opportunity for reporting bias by enabling comparison of the completed review with what was planned in the protocol. PROSPERO, therefore, provides access to ongoing reviews as well as completed and / or published reviews.
Epistemonikos is a web-based bibliographic service which provides access to many thousands of systematic reviews, broad syntheses of reviews and structured summaries, and their included primary studies (http://www.epistemonikos.org/en). The aim of Epistemonikos is to provide rapid access to systematic reviews in health. Epistemonikos uses the eligibility criteria specified by the review authors to include primary studies in the database. Records that are classified as systematic reviews within Epistemonikos are now available through the Cochrane Library but are only included in search results for queries entered in the Basic Search box, available from the Cochrane Library header. They are not retrieved when using Advanced Search.
The Systematic Review Data Repository (SRDR) is an open and searchable archive of systematic reviews and their data (http://srdr.ahrq.gov/).
Health Systems Evidence is a repository of evidence syntheses about governance, financial and delivery arrangements within health systems, and about implementation strategies that can support change in health systems. The types of syntheses include evidence briefs for policy, overviews of systematic reviews, systematic reviews, protocols, and registered titles. The audience is policy makers / researchers (https://www.healthsystemsevidence.org).
Specific evidence-based search services such as Turning Research into Practice (TRIP) (https://www.tripdatabase.com/) can also be used to identify reviews and guidelines (Brassey 2007). For the range of systematic review sources searched by TRIP see www.tripdatabase.com/about. Access is offered at two levels: free of charge and subscription.
SUMSearch 2 (http://sumsearch.org/) simultaneously searches for original studies, systematic reviews, and practice guidelines from multiple sources.
MEDLINE, Embase and other bibliographic databases, such as CINAHL (Wright et al 2015), can also be used to identify review articles and guidelines. For the 2019 release of the Medical Subject Headings (MeSH), Systematic Review was introduced as a Publication Type term. NLM announced: “We added the publication type ‘Systematic Review’ retrospectively to appropriate existing MEDLINE citations. With this re-indexing, you can retrieve all MEDLINE citations for systematic reviews and identify systematic reviews with high precision.”
https://www.nlm.nih.gov/pubs/techbull/ma19/brief/ma19_systematic_review.html
Embase has a thesaurus (Emtree) term ‘Systematic Review’, which was introduced in 2003. For records prior to 2003, the Emtree terms ‘review’ or ‘evidence-based medicine’ could be used.
Several filters to identify reviews and overviews of systematic reviews in MEDLINE (Boynton et al 1998, Glanville et al 2001, Montori et al 2005, Wilczynski and Haynes 2009) and Embase have been developed and tested over the years (Wilczynski et al 2007, Lunny et al 2015). Until late 2018, the PubMed Systematic Reviews filter under the Clinical Queries link was very broad in its scope and retrieved many references that were not systematic reviews. The strategy was defined by NLM as follows: “This strategy is intended to retrieve citations identified as systematic reviews, meta-analyses, reviews of clinical trials, evidence-based medicine, consensus development conferences, guidelines, and citations to articles from journals specializing in review studies of value to clinicians. This filter can be used in a search as systematic [sb].” An archived version of this search filter is available from the InterTASC Information Specialists’ Sub-Group’s Search Filter Resource at:
This search filter was replaced by NLM in late 2018 with a much more precise filter and is defined by NLM as follows: “This strategy is intended to retrieve citations to systematic reviews in PubMed and encompasses: citations assigned the ‘Systematic Review’ publication type during MEDLINE indexing; citations that have not yet completed MEDLINE indexing; and non-MEDLINE citations. This filter can be used in a search as systematic [sb].”
Example: exercise hypertension AND systematic [sb]
This filter is also available on the Filters sidebar under ‘Article types’ and on the Clinical Queries screen. The full search filter is available at:
https://www.nlm.nih.gov/bsd/pubmed_subsets/sysreviews_strategy.html
The sensitive Clinical Queries Filters for therapy, diagnosis, prognosis, and aetiology perform well in retrieving not only primary studies but also systematic reviews in PubMed. In a test of the Clinical Queries Filters by the McMaster Health Information Research Unit (HIRU), Wilczynski and colleagues reported that performance could be improved by combining the Clinical Queries Filters with the HIRU systematic review filter using the Boolean operator ‘OR’ (Wilczynski et al 2011). As well as filters for study design, some filters are available for special populations, and these might be combined with systematic review filters (Boluyt et al 2008).
Research has been conducted to help researchers choose the filter appropriate to their needs (Lee et al 2012, Rathbone et al 2016). Filters and current reviews of filter performance can be found on the InterTASC Information Specialists’ Subgroup Search Filter Resource website (https://sites.google.com/a/york.ac.uk/issg-search-filters-resource/filters-to-identify-systematic-reviews) (Glanville et al 2019a). For further information on search filters see Section 3.6 and subsections.
National and regional drug approval and reimbursement agencies may also be useful sources of reviews:
- The Agency for Healthcare Research and Quality (AHRQ) publishes systematic reviews and meta-analyses. Evidence reports, comparative effectiveness reviews, technical briefs, Technology Assessment Program reports, and U.S. Preventive Services Task Force evidence syntheses are available under the Evidence-based Practice Centers (EPC) Program of the Agency for Healthcare Research and Quality. Access to the evidence reports is provided at: http://www.ahrq.gov/research/findings/evidence-based-reports/search.html.
- The Canadian Agency for Drugs and Technologies in Health (CADTH) (www.cadth.ca) is an independent, not-for-profit organization responsible for providing healthcare decision-makers with evidence reports to help make informed decisions about the optimal use of drugs, diagnostic tests, and medical, dental, and surgical devices and procedures. CADTH’s Common Drug Review reports, Pan Canadian Oncology Drug Review reports, Health Technology Assessments, Technology Reviews and Therapeutic Reviews are published in full text on their website and include the full search strategy for the clinical evidence used in that review.
- The National Institute for Health and Care Excellence (NICE) (www.nice.org.uk) publishes guidance that includes recommendations on the use of new and existing medicines and other treatments within the National Health Service (NHS) in England and Wales. These reviews can be about medicines, medical devices, diagnostic tests, surgical procedures, or health promotion activities. Each guidance and appraisal document is based on a review of the evidence and reports the searches used.
Clinical guidelines, based on reviews of evidence, may also provide useful information about the search strategies used in their development: see the Appendix for examples of sources of clinical guidelines. Guidelines can also be identified by searching MEDLINE where guidelines should be indexed under the Publication Type term ‘Practice Guideline’, which was introduced in 1991. Embase has a thesaurus term ‘Practice Guideline’, which was introduced in 1994.
The ECRI Guidelines Trust (https://guidelines.ecri.org/) provides access to a free web-based repository of objective, evidence-based clinical practice guideline content. It includes evidence-based guidance developed by nationally and internationally recognized medical organizations and medical specialty societies. Guidelines are summarized and appraised against the US Institute of Medicine (IOM) Standards for Trustworthiness. The Guidelines Trust provides the following guideline-related content:
- Guideline Briefs: summarizes content providing the key elements of the clinical practice guideline.
- TRUST (Transparency and Rigor Using Standards of Trustworthiness) Scorecards: ratings of how well guidelines fulfil the IOM Standards for Trustworthiness.
The Agency for Healthcare Research and Quality (AHRQ)’s National Guideline Clearinghouse existed as a public resource for summaries of evidence-based clinical practice guidelines but ceased production in July 2018 with the latest guidelines being accepted for inclusion until March 2018. The resource offered systematic comparisons of selected guidelines that addressed similar topic areas. For further information as to whether this resource will be reintroduced see: https://www.ahrq.gov/gam/updates/index.html.
Evidence summaries such as online / electronic textbooks, point-of-care tools and clinical decision support resources are a type of synthesized medical evidence. Examples of these tools include BMJ Clinical Evidence, ClinicalKey, DynaMed Plus and UpToDate in addition to Cochrane’s own point-of-care tool Cochrane Clinical Answers. Although they are designed to be used in clinical practice, they offer evidence for diagnosis and treatment of specific conditions and are regularly updated with links to and reference lists to reports of relevant studies which can help in identifying studies, reviews, and overviews. Most evidence summaries for use in clinical practice are available via subscription to commercial vendors.
As noted above, it is mandatory, for authors of Cochrane reviews of interventions, to check reference lists of included studies and any relevant systematic reviews identified (MECIR C30). Checking reference lists within eligible studies supplements other searching approaches and may reveal new studies, or confirm that the topic has been thoroughly searched (Greenhalgh and Peacock 2005, Horsley et al 2011). Examples of situations where checking reference lists might be particularly beneficial are:
- when the review is of a new technology;
- when there have been innovations to an existing technique or surgical approach;
- where the terminology for a condition or intervention has evolved over time; and
- where the intervention is one which crosses subject disciplines, for example, between health and other fields such as education, psychology or social work. Researchers may use different terminology to describe an intervention depending on their field (O'Mara-Eves et al 2014).
It is not possible to give overall guidance as to which of the above sources should be searched in the case of all reviews to identify other reviews, guidelines and reference lists as sources of studies. This will vary from review to review. Reviews authors should discuss this with their Cochrane Information Specialist or their medical / healthcare librarian or information specialist.
1.3.5 General web searching (including search engines / Google Scholar etc)
Searching the World Wide Web (hereafter, web) involves using resources which are not specifically designed to host and facilitate the identification of studies. This includes general search engines such as Google Search and the websites of organizations that are topically relevant for review topics, such as charities, research funders, manufacturers and medical societies. These resources often have basic search interfaces and host a wide range of content, which poses challenges when conducting systematic searching (Stansfield et al 2016). Despite these challenges web searching has the potential to identify studies that are eligible for inclusion in a review, including ‘unique’ studies that are not identified by other search methods (Eysenbach et al 2001, Ogilvie et al 2005, Stansfield et al 2014, Godin et al 2015, Bramer et al 2017a). It is good practice to carry out web searching for review topics where studies are published in journals that are not indexed in bibliographic databases or where grey literature is an important source of data (Ogilvie et al 2005, Stansfield et al 2014, Godin et al 2015). Grey literature is literature “which is produced on all levels of government, academics, business and industry in print and electronic formats, but which is not controlled by commercial publishers” (see Section 1.1.6) (Farace and Frantzen 1997, Farace and Frantzen 2005).
It is good practice to base the search terms used for web searching on the search terms used for searching bibliographic databases (Eysenbach et al 2001). A simplified approach, however, might be required due to the basic search interfaces of web resources. For example, web resources are unlikely to support multi-line search strategy development or nested use of Boolean operators, and single-line searching is often limited by a maximum number of alphanumeric characters. As such, it might be necessary to rewrite a search using fewer search terms or to conduct several searches of the same resource using different combinations of search terms (Eysenbach et al 2001, Stansfield et al 2016). In addition to using search terms, web searching involves following links to webpages and websites. This is less structured than searching using pre-specified search terms and the searcher will need to use their discretion to decide when to start and stop searching (Stansfield et al 2016). Wherever possible, a similar approach to searching should be used for different web resources to ensure consistency and searches should be documented in full and reported in the review (see Chapter 4, Section 4.5).
Web resources are unlikely to have a function for exporting results to reference management software, in which case the searcher may decide to screen the results ‘on screen’ while searching. Alternatively, screenshots can be taken and screened at a later time (Stansfield et al 2016). This process can be facilitated by software such as Evernote or OneNote. Because website content can be deleted or edited by the website editor at any time, a permanent record of any relevant studies should be retained.
Web searching should use a combination of search engines and websites to ensure a wide range of sources are identified and searched in depth.
Search engines
Due to the scale and diversity of content on the web, searching using a search engine is likely to retrieve an unmanageable number of results (Mahood et al 2014). Results are usually ranked according to relevance as determined by a search engine’s algorithm, so it might be useful to limit the screening process to a pre-specified number of results, e.g. limits ranging from 100 to 500 results have been reported in recent Cochrane Reviews (Briscoe 2018). Alternatively, an ad hoc decision to stop screening can be made when the search results become less relevant (Stansfield et al 2016). It is good practice to use a more comprehensive approach when screening Google Scholar results, which are limited to 1000, to ensure that all relevant studies, including grey literature, are identified (Haddaway et al 2015). Some search engines allow the user to limit searches to a specified domain name or file type, or to web pages where the search terms appear in the title. These options might improve the precision of a search though they might also reduce its sensitivity. The reported number of results identified by a search engine is usually an estimate which varies over time, and the actual number of results might be much lower than reported (Bramer 2016). Search engines often combine search terms using the ‘AND’ Boolean operator by default. Some search engines support additional search operators and features such as ‘OR’, ‘NOT’, wildcards and phrase searching using quotation marks.
There are many freely available search engines, each of which offers a different approach to searching the web. Because each search engine uses a different algorithm to retrieve and rank its results, the results will differ depending on the search engine that is used (Dogpile.com 2007). Some search engines use internet protocol (IP) addresses to tailor the search results to a user’s search history, so the search results might differ between users. For these reasons, it might be worth experimenting with or combining different search engines to retrieve a wider selection of results. There are freely available meta-search engines which search a combination of search engines, though they are often limited with regard to which search engines can be combined.
A selection of freely available search engines and meta-search engines is shown in Box 1.a.These are examples of different types of search engine rather than a list of recommended search engines. No specific search engines are recommended for a Cochrane Review.
Box 1.a Search engines
|
Dogpile http://www.dogpile.com/ Dogpile is a meta-search engine which in a study from 2007 is reported to search Google Search, Yahoo!, Ask and Bing (Dogpile.com 2007). A more up to date list of search engines used by Dogpile has not been identified. DuckDuckGo https://duckduckgo.com/ DuckDuckGo protects the privacy of its users by not recording their IP addresses and search histories. A potential advantage for systematic review authors is that DuckDuckGo does not use search histories to personalize its search results, which might make it better at ranking less frequently visited but useful pages higher in the results. Google Scholar https://scholar.google.com/ Google Scholar is a specialized version of Google Search which limits results to scholarly literature, including published studies and grey literature. It cannot be used instead of searching bibliographic databases due to its basic search interface and a block on viewing more than 1000 records per search (Boeker et al 2013a, Bramer et al 2016a). It can, however, be a useful resource when used alongside bibliographic databases for identifying studies and grey literature not indexed in bibliographic databases or not retrieved by the bibliographic database search strategies (Haddaway et al 2015, Bramer et al 2017a). The option to search the full text of studies can contribute to the identification of unique studies when using similar or the same search terms as used in bibliographic databases (Bramer et al 2017a). References can be exported to reference management software, though the number of references that can be exported at a time is limited to 20 (Bramer et al 2013). Google Search https://www.google.com/ Google Search is the most widely used search engine worldwide. An advantage of its popularity is that there is an abundance of online material on how to make the most of its advanced search features. The Verbatim feature in the Google Search Tools menu can be used to ensure search results contain the precise search terms used (e.g. will not retrieve “nursing” if searching for “nurse”) and to switch off the personalization of search results based on websites which the user has previously visited. Personalization can also be deactivated via the settings menu. Microsoft Academic https://academic.microsoft.com/ Microsoft Academic is a scholarly search engine which, like Google Scholar, indexes scholarly literature. It was relaunched in 2016 after a four year hiatus. Comparative studies of Google Scholar and Microsoft Academic show that Google Scholar indexes more content than Microsoft Academic (Gusenbauer 2019). Microsoft Academic, however, has more structured and richer metadata than Google Scholar, which is reported to facilitate better search functionality and handling of results (Hug et al 2017). |
Not all content on websites is indexed by search engines, so it is important to consider accessing and searching any potentially useful websites which are identified in the results (Devine and Egger-Sider 2013).
Websites
The selection of websites to search will be determined by the review topic. It is good practice to investigate whether the websites of relevant pharmaceutical companies and medical device manufacturers host trials registers which should be searched for studies. The websites of medicines regulatory bodies such as the US Food and Drug Administration (FDA) and the European Medicines Agency (EMA) should be searched for regulatory documentation (see Section 1.2 and subsections). It might also be useful to search the websites of professional societies, national and regional health departments, and health related non-governmental organizations and charities for studies not indexed in bibliographic databases and grey literature (Ogilvie et al 2005, Godin et al 2015).
Searching websites will usually yield a lower number of results than search engines, so it should be possible to screen all the results rather than a pre-specified number.
1.4 Summary points
- Cochrane Review authors should seek advice from their Cochrane Information Specialist on sources to search.
- Authors of non-Cochrane reviews should seek advice from their medical / healthcare librarian or information specialist, with experience of conducting searches for studies for systematic reviews.
- The key database sources which should be searched are the Cochrane Review Group’s Specialized Register (internally, e.g. via the Cochrane Register of Studies, or externally via CENTRAL), CENTRAL, MEDLINE and Embase (if access to Embase is available to either the review authors or the CRG).
- Appropriate national, regional and subject specific bibliographic databases should be searched according to the topic of the review.
- Relevant grey literature sources such as those containing reports, dissertations/theses and conference abstracts should be searched.
- Searches should be conducted to locate previous reviews on the same topic, to identify additional studies included in (and excluded from) those reviews.
- Reference lists of included studies should be checked to identify additional studies.
- Trials registers and repositories of results, such as regulatory agency sources, where relevant to the topic, should be searched through both ClinicalTrials.gov and the WHO International Clinical Trials Registry Platform (ICTRP) portal and other sources as appropriate.
- Regulatory agency sources and clinical study reports should also be considered as sources for study data.
- Citation indexes should be considered as an additional source of relevant studies.
2 Planning the search process
2.1 Cochrane-wide search initiatives and the Cochrane Centralized Search Service
It is unlikely that CENTRAL will ever contain all reports of randomized trials. Substantial efforts are, however, underway to populate this unique resource with as many reports as possible in a systematic, transparent and efficient way so as to help information specialists and systematic review authors find relevant evidence quickly and reliably. Given that CENTRAL will likely never be 100% comprehensive, searching across other major databases will remain a core activity for the foreseeable future.
Information specialists should consider numerous factors when deciding which sources to include in their searches. These include: being aware of the time taken for records to appear in CENTRAL from source databases such as MEDLINE and Embase, understanding that across the years different processes and searches have been used to populate CENTRAL, and recognizing that for trial registry records not all fields of content available for those records in their source databases are included in CENTRAL. Work is underway to assess the comprehensiveness of CENTRAL in order to be able to provide users of CENTRAL with as much information as possible regarding the need to search beyond CENTRAL for RCT evidence.
New processes in the form of crowdsourcing and machine learning are increasingly being used to help populate CENTRAL in addition to ‘direct feeds’ of records.
Table 2.1.a is designed to be a quick reference to current sources that feed into CENTRAL.
Table 2.1.a Sources searched as part of the Cochrane Centralized Search Service (CSS)
|
Source |
Process type |
Detail |
Current schedule |
|
MEDLINE (searched via PubMed) (see also under Embase below) |
Direct feed based on index terms |
Records indexed as RCT or CCT publication type (all dates) More details see Section 2.1.1 |
Monthly feed. New records appear in CENTRAL during 3rd week of every month |
|
Embase (searched via Embase.com - including ‘native MEDLINE’ records) |
Direct feed based on index term |
Records indexed as RCT Emtree term (all dates) More details see Section 2.1.2 |
Monthly feed. New records appear in CENTRAL during 3rd week of every month |
|
Direct feed based on index term |
Records indexed as CCT Emtree term (2010 to Dec 2017) More details see Section: 2.1.2 |
This was a monthly feed. New records appeared in CENTRAL during 3rd week of every month. It was stopped at the start of 2018 due to the number of records indexed as CCT that were found not to be randomized or quasi-randomized trials |
|
|
Cochrane Crowd and machine learning RCT Classifier |
The results retrieved from a sensitive search in Embase performed every month are put through a specially developed machine learning RCT Classifier. Based on scores assigned, some records are rejected at that stage, while the rest go to Cochrane Crowd for assessment More details see Section: 2.1.2 |
The searches are run monthly. The Cochrane Crowd varies in how quickly those results are screened. Allow two months for records to be screened and resolved where necessary. New records appear in CENTRAL during 3rd week of every month |
|
|
ClinicalTrials. |
Direct feed based on ClinicalTrials. |
ClinicalTrials.gov records across all dates to March 2018 which received an RCT Classifier score of 80% or more were submitted to CENTRAL in March 2018 More details see Section: 2.1.3.2 |
From April 2018, a monthly feed of ClinicalTrials.gov records with an RCT Classifier score of 80% or more continues to be fed into CENTRAL. New records appear in CENTRAL during 3rd week of every month |
|
Crowdsourced feed |
ClinicalTrials.gov records with a classifier score of below 80% are assessed by Cochrane Crowd More details see Section: 2.1.3.2 |
From April 2018, the backlog of records from this source was cleared and submitted to CENTRAL. From then on new records are added each month during the 3rd week |
|
|
International Clinical Trials Registry Platform (ICTRP) |
Direct feed based on search query made on the XML export |
((randomised OR randomized) NOT (randomised: no OR randomized: no)) (see footnote beneath Table) More details see Section 2.1.3.3 |
Records were added to CENTRAL in March 2019 Thereafter new records meeting the direct feed criteria are added to CENTRAL each month during the 3rd week |
|
Crowdsourced feed |
Records that did not meet the direct feed criteria were sent to Cochrane Crowd More details see Section 2.1.3.3 |
Records were added to CENTRAL in March 2019 Thereafter, new records not meeting the direct feed criteria but identified by the Crowd as RCTs are added to CENTRAL each month during the 3rd week |
|
|
KoreaMed |
Manual screening (all dates to July 2017) |
Using a sensitive search strategy, records sourced from KoreaMed were screened across all dates to January 2014. From January 2014, all KoreaMed records were manually screened up to July 2017 More details see Section: 2.1.3.4 |
Records added to CENTRAL in 2017 |
|
Cochrane Crowd and machine learning RCT classifier (from August 2017) |
KoreaMed records with a classifier score of above 10% are assessed by Cochrane Crowd More details see Section: 2.1.3.4 |
Since August 2017, records that receive a score of 10% or less are automatically rejected. Records that receive a score of 11% or above are sent to Cochrane Crowd for screening |
Footnote: ‘no’ in the International Clinical Trials Registry Platform (ICTRP) entry above refers to the picklist value selected by those registering their trial in ICTRP to indicate that the trial is not a randomized controlled trial. Records where the picklist value was ‘no’ in answer to this question about study design were excluded from the set of records directly fed into CENTRAL. Instead they were manually screened. Figure 2.a illustrates the contents of CENTRAL.
Figure 2.a Illustration of the contents of CENTRAL
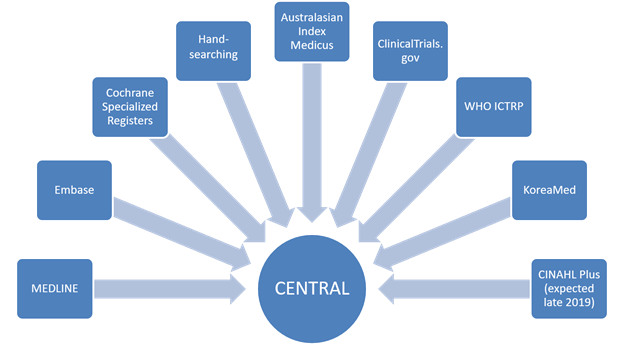
In 2015, building on the processes established for the Embase project, to identify records from Embase and MEDLINE (see Section 2.1.2), Cochrane began a pilot initiative with the objective of adding to the number of sources to be searched and screened ‘centrally’, known as the Cochrane Centralized Search Service (CSS). The CSS initiative is still underway at the time of writing (August 2019). There are currently five databases searched as part of the CSS. They are MEDLINE / PubMed (see Section 2.1.1), Embase (see Section 2.1.2), ClinicalTrials.gov (see Section 2.1.3.2), the WHO International Clinical Trials Registry Platform (ICTRP) (see Section 2.1.3.3) and KoreaMed (see Section 2.1.3.4). In late 2019, it is expected that CINAHL Plus (EBSCOhost) (see Section 2.1.3.5) will become the sixth source to be searched and screened for reports of randomized trials as part of the Centralized Search Service. All sources are searched or queried via an API each month. Where possible, no filters or limits are applied in an effort to achieve maximum sensitivity. For both Embase and CINAHL Plus, however, a methodological filter has been developed for each source.
Each of the CSS sources had ‘backlogs’ to deal with in parallel to setting up prospective routines to identify newly indexed reports of RCTs. The backlogs for Embase, ClinicalTrials.gov and ICTRP have all been cleared. This was achieved by using a combination of machine learning in the form of the RCT classifier and crowdsourcing via Cochrane Crowd. The CSS aims to provide systematic review authors and others with an even baseline of access to the relevant evidence needed to produce systematic reviews and other evidence products. It is unlikely it will ever completely replace the need for multi-source, bespoke, review-based searches, especially for cross-disciplinary or complex reviews, but it is hoped that it will substantially improve access to RCT evidence and reduce the amount of multi-source searching currently needed. A retrospective analysis is currently underway (August 2019) to evaluate the performance of the CSS and to identify any potential areas for improvement. The results of this analysis will be presented at the 26th Cochrane Colloquium in 2019 (Noel-Storr et al 2019).
2.1.1 What is in the Cochrane Central Register of Controlled Trials (CENTRAL) from MEDLINE?
CENTRAL contains all records from MEDLINE indexed with the Publication Type term ‘Randomized Controlled Trial’ or ‘Controlled Clinical Trial’ except those that are indexed solely as animal studies (not also as human studies). For further details see the CENTRAL Creation Details file in the Cochrane Library:
https://www.cochranelibrary.com/central/central-creation
A substantial proportion of the MEDLINE records coded ‘Randomized Controlled Trial’ or ‘Controlled Clinical Trial’ in the Publication Type field have been coded as a result of the work within Cochrane (Dickersin et al 2002). Handsearch results from Cochrane entities, for journals indexed in MEDLINE, were sent to the US National Library of Medicine (NLM), where the MEDLINE records were re-tagged with the publication types ‘Randomized Controlled Trial’ or ‘Controlled Clinical Trial’ as appropriate. In addition, the US Cochrane Center (formerly the New England Cochrane Center, Providence Office and the Baltimore Cochrane Center and now Cochrane US) and the UK Cochrane Centre (now Cochrane UK) conducted an electronic search of MEDLINE from 1966 to 2004 to identify reports of randomized trials, identifiable from the MEDLINE titles and / or abstracts, not already indexed as such, using the first two phases of the Cochrane Highly Sensitive Search Strategy first published in 1994 (Dickersin et al 1994) and thereafter updated and included in subsequent editions of this Handbook. The free-text terms used were: clinical trial; (singl$ OR doubl$ OR trebl$ OR tripl$) AND (mask$ OR blind$); placebo$; random$. The $ sign indicates the use of a truncation symbol. The subject heading terms (MeSH) used were (‘exploded’ where possible to include narrower, more specific terms): randomized controlled trials; random allocation; double-blind method; single-blind method; clinical trials; placebos. The following subject heading term (MeSH) was used ‘unexploded’: research design. The Publication Type terms used were: randomized controlled trial; controlled clinical trial; clinical trial.
A test was carried out using the terms in phase three of the 1994 Cochrane Highly Sensitive Search Strategy but the precision of those terms, having already searched on all the terms in phases one and two as listed above, was considered to be too low to warrant using these terms for the above project (Lefebvre and Clarke 2001). It was, however, recognized that some of these terms might be useful when combined with subject terms to identify studies for some specific reviews (Eisinga et al 2007).
The above search was limited to humans. The following years were completed by the US Cochrane Center (1966 to 1984; 1998 to 2004) and by the UK Cochrane Centre (1985 to 1997). The results for these years were forwarded to the NLM and re-tagged in MEDLINE and are thus included in CENTRAL. More recent MEDLINE records, which are now included, under licence, in Embase, are being searched as part of the Embase screening project (see Section 2.1.2).
CENTRAL includes from MEDLINE not only reports of trials that meet the more restrictive Cochrane definition for a quasi-randomized trial (indexed in MEDLINE as ‘Controlled Clinical Trial’) (Box 2.a) but also trial reports that meet the less restrictive NLM definition (Box 2.b) which includes historical comparisons. There is currently no method of distinguishing, either in CENTRAL or in MEDLINE, which of these records meet the more restrictive Cochrane definition, as they are all indexed with the Publication Type term ‘Controlled Clinical Trial’.
Box 2.a Cochrane definitions and criteria for randomized controlled trials (RCTs) and quasi-randomized trials
|
Records identified for inclusion should meet the eligibility criteria devised and agreed in November 1992, which were first published, in 1994, in the first version of this Handbook (Oxman et al 1994). According to these eligibility criteria: A trial is eligible if, on the basis of the best available information (usually from one or more published reports), it is judged that:
Trials eligible for inclusion are classified according to the reader’s degree of certainty that random allocation was used to form the comparison groups in the trial. If the author(s) state explicitly (usually by some variant of the term ‘random’ to describe the allocation procedure used) that the groups compared in the trial were established by random allocation, then the trial is classified as a RCT (randomized controlled trial). If the author(s) do not state explicitly that the trial was randomized, but randomization cannot be ruled out, the report is classified as a CCT (controlled clinical trial). The classification CCT is also applied to quasi-randomized studies, where the method of allocation is known but is not considered strictly random, and also trials that are possibly quasi-randomized. Examples of quasi-random methods of assignment include alternation, date of birth, and medical record number. The classification as RCT or CCT is based solely on what the author has written, not on the reader’s interpretation; thus, it is not meant to reflect an assessment of the true nature or quality of the allocation procedure. For example, although ‘double-blind’ trials are nearly always randomized, many trial reports fail to mention random allocation explicitly and should therefore be classified as CCT. Relevant reports are reports published in any year, of studies comparing at least two forms of health care (healthcare treatment, healthcare education, diagnostic tests or techniques, a preventive intervention, etc) where the study is on either living humans or parts of their body or human parts that will be replaced in living humans (e.g. donor kidneys). Studies on cadavers, extracted teeth, cell lines, etc are not relevant. Searchers should identify all controlled trials meeting these criteria regardless of relevance to the entity with which they are affiliated. The highest possible proportion of all reports of controlled trials of health care should be included in CENTRAL. Thus, those searching the literature to identify trials should give reports the benefit of any doubts. Review authors will decide whether to include a particular report in a review. |
In 2013, a Cochrane working group was formed to review the record type eligibility for CENTRAL and to ensure consistency of practice and guidance for the Embase project and handsearcher training. This group focused on types of report rather than types of study. The group determined that reports of protocols for randomized or quasi-randomized trials, along with letters, replies, errata, and retractions relating to RCTs or quasi-RCTs are all to be included in CENTRAL.
Box 2.b US National Library of Medicine 2019 definitions (Scope Notes) for the Publication Type terms ‘Randomized Controlled Trial’ and ‘Controlled Clinical Trial’
Randomized Controlled TrialA work that reports on a clinical trial that involves at least one test treatment and one control treatment, concurrent enrollment and follow-up of the test- and control-treated groups, and in which the treatments to be administered are selected by a random process, such as the use of a random-numbers table. Controlled Clinical TrialA work that reports on a clinical trial involving one or more test treatments, at least one control treatment, specified outcome measures for evaluating the studied intervention, and a bias-free method for assigning patients to the test treatment. The treatment may be drugs, devices, or procedures studied for diagnostic, therapeutic, or prophylactic effectiveness. Control measures include placebos, active medicine, no-treatment, dosage forms and regimens, historical comparisons, etc. When randomization using mathematical techniques, such as the use of a random numbers table, is employed to assign patients to test or control treatments, the trial is characterized as a RANDOMIZED CONTROLLED TRIAL. |
MEDLINE records are also currently being added into CENTRAL from Embase. Since 2010, Elsevier has included MEDLINE records in Embase under licence with the US National Library of Medicine (see further details in Section 2.2.2 on specific issues when searching Embase).
2.1.2 What is in the Cochrane Central Register of Controlled Trials (CENTRAL) from Embase?
A retrospective search conducted by the UK Cochrane Centre (now Cochrane UK) for reports of trials in Embase has been undertaken for the years 1974 to 2010. For the years 1974 to 1979, the free-text terms: random$; factorial$; crossover$; cross-over$; and placebo$ were used. For the years 1980 to 2008, the following free-text terms: random$; factorial$; crossover$; cross-over$; cross over$; placebo$; doubl$ adj blind$; singl$ adj blind$; assign$; allocat$; volunteer$; and the following index terms, known as Emtree terms: crossover-procedure; double-blind procedure; randomized controlled trial; single-blind procedure were used. For 2009, the following free-text terms: random$; crossover$; cross-over$; cross over$; placebo$; doubl$ adj blind$; singl$ adj blind$; allocat$; and the following index terms, known as Emtree terms: crossover-procedure; double-blind procedure; randomized controlled trial; single-blind procedure were used. In addition, the following terms were searched limited to the title only: trial, comparison. For 2010, the following free-text terms were searched limited to the title, abstract and original title fields only: crossover$, cross over$, placebo$, doubl$ adj blind$, allocat$, random$; and limited to the title only: trial; and the following index terms were searched: crossover-procedure, double-blind procedure, single-blind procedure and randomized controlled trial. (Note: cross over$ includes cross-over$ in Ovid syntax).
The searches across all years of this project (1974 to 2010) yielded a total of approximately 100,000 reports of trials not indexed, at the time of the search, as randomized controlled trial or controlled clinical trial in MEDLINE. All of these reports are now published in CENTRAL (Lefebvre et al 2008). The final submission of reports under this project, of trials identified in journal article records added to Embase in 2010, was published in CENTRAL in February 2012. This project then formally ended, with a newly funded project starting in 2013.
In March 2013, Cochrane launched a further Embase Project to provide ongoing screening of records from Embase to identify additional reports of trials. This project was co-ordinated by Metaxis Ltd., the Cochrane Dementia and Cognitive Improvement Group and York Health Economics Consortium. Initially, a search covering January 2011 to December 2013, inclusive, was run, from which 28,442 unique Embase records were identified and published in CENTRAL, January 2014 (Issue 1). All these records were identified from a search in Embase (via Ovid) using the Emtree headings Randomized Controlled Trial (RCT) or Controlled Clinical Trial (CCT). It is estimated that this search, using only these two headings, identified two-thirds of records eligible for inclusion in CENTRAL from the 2011 to 2013 period.
The remaining records were identified using the search strategy developed by the UK Cochrane Centre, described above, with records indexed as either RCT or CCT removed, as those records had already been identified and added to CENTRAL. A small team of expert screeners screened the results retrieved and identified a further 20,655 records eligible for CENTRAL.
In parallel to the work described above, a new search filter to identify potential reports of randomized trials in Embase was developed in 2013 and initiated in January 2014. It was developed following an examination of 1000 relevant reports (reference standard) of randomized trials, and was tested on a second set of 1000 records. The filter was tiered. The first tier identified records with the most relevant EMTREE headings RANDOMIZED CONTROLLED TRIAL or CONTROLLED CLINICAL STUDY. The second tier comprised search terms likely to find records from the reference standard which did not contain those two EMTREE headings (http://www.cochranelibrary.com/help/central-creation-details.html). The filter was amended in the light of information gained from screening and was revised to minimize false negatives. The revised filter was used from January 2015 and the second tier now includes a series of search terms (study design and animal experiment terms), which are excluded from the results. In September 2017 the filter was amended once again by removing the term CONTROLLED CLINICAL STUDY from the tier 1 search and adding it to the tier 2 search. This was done because it was felt that the CONTROLLED CLINICAL STUDY term was adding too many false positives directly into CENTRAL. Adding the term to the tier 2 search means that these records now go through Cochrane Crowd.
Records are screened using a crowdsourcing model, accessible from the Cochrane Crowd micro-tasking platform http://crowd.cochrane.org/index.html. Here, Cochrane contributors and members of the general public can contribute to screening records from Embase after completing a brief training exercise. As at May 2019 over 550,000 records had been collectively screened, and over 52,000 additional reports of trials had been identified and added to CENTRAL.
In 2009, Elsevier began adding conference records to Embase, and to date (August 2019) has added about 2.5 million conference abstracts from about 7000 conferences (https://www.elsevier.com/solutions/embase-biomedical-research/embase-coverage-and-content). This created a sizable backlog of records. The Embase screening project searched and downloaded all records (not just conference abstracts) added to Embase between 2010 and 2013 inclusive. The search strategy used for the conference ‘backlog’ was the most recent version in use by the UK Cochrane Centre. This was so that screening of this backlog could get underway quickly whilst the new search filter was being developed. All reports of RCTs identified from the screening of these records had been published in CENTRAL by the end of 2014.
Introducing machine learning into the workflow
In January 2016 the machine learning RCT Classifier was used for the first time on records identified from Embase via the monthly sensitive search described above. Records that received a likelihood score below a pre-specified cut-off-point were deemed to be not RCTs and no further action was taken on them. Those records that scored on or above the cut-off-point were then sent to Cochrane Crowd for manual assessment. This has remained the workflow for Embase records since the start of 2016. Work to evaluate the potential and the performance of the RCT classifier can be found in (Wallace et al 2017) and (Marshall et al 2018). In terms of the application of the RCT classifier to the central feed of Embase records, approximately 50% of records score below the currently used cut-off-point representing a significant reduction in manual screening required by the Crowd. (See Chapter 4, Section 4.6.6.2 for further information about using machine learning to classify reports of RCTs.)
2.1.3 What is in the Cochrane Central Register of Controlled Trials (CENTRAL) from other non-Cochrane sources and handsearching?
2.1.3.1 Introduction
Many CRGs and Fields have undertaken searching of the specialist healthcare literature (both journals and databases) in their areas of interest. More than 3000 journals have been, or are being, handsearched. Identified trial reports that are not relevant to a CRG’s scope and thus are not appropriate for their Specialized Register (see Section 2.1.4) are published in CENTRAL as handsearch results. Handsearch records can be identified in CENTRAL as they are assigned the tag HS-HANDSRCH in addition to a source code indicating the Centre, Field or Review Group that submitted the record (see https://www.cochranelibrary.com/central/central-creation)
The Australasian Cochrane Centre (now Cochrane Australia) co-ordinated a search of the National Library of Australia’s Australasian Medical Index from 1966 (McDonald 2002). This search was updated to include records added up to December 2009, when the database ceased to be updated (it is now available as an archived database from RMIT Publishing (https://www.informit.org/index-product-details/AMI). All records identified have been added to CENTRAL.
The Chinese Cochrane Center (now Cochrane China), with support from the Australasian Cochrane Centre (now Cochrane Australia), the UK Cochrane Centre (now Cochrane UK) and Cochrane centrally has co-ordinated a search of the Chinese Biomedical Literature Database (CBM) from 1978 to 2008 and has identified approximately 30,000 reports of trials. These records have not been added to CENTRAL.
2.1.3.2 Records from ClinicalTrials.gov
From August 2017, eligible ClinicalTrials.gov (CT.gov) (https://clinicaltrials.gov/) records are being identified and systematically added to CENTRAL through Cochrane’s Centralized Search Service project and Project Transform.
Process description
All CT.gov records will go through Cochrane’s RCT machine classifier and some go through Cochrane Crowd (crowd.cochrane.org). The classifier provides likelihood scores for each record being either a randomized or quasi-randomized trial report. Records with an 80% or greater likelihood score will be submitted directly to CENTRAL. Records with a 10% or less likelihood score will be rejected without any further action. Records with a likelihood score of 11% to 79% will be sent to Cochrane Crowd to be screened by humans. Performance evaluations show over 99% accuracy at the thresholds described above.
Backlog
In September 2017, the date at which the CSS initiative began to process records from ClinicalTrials.gov centrally, ClinicalTrials.gov contained approximately 250,000 records. This is what the CSS project team termed the ClinicalTrials.gov backlog. Of those, 72,030 records had an RCT Classifier score of 10% or less; these records were rejected. 74,801 had a score of 80% or more; these records were de-duplicated against CENTRAL and unique records were added to CENTRAL in April 2018 (available in issue 4).
The remaining 102,097 records with a likelihood score of between 11% and 79% were sent to Cochrane Crowd. This backlog was cleared by the end of April 2018. The records were added to CENTRAL in May 2018.
Field mappings
The CT.gov records contain several fields, but not all fields are included in CENTRAL. The fields that are displayed in CENTRAL are the Public and Scientific titles, the URL to the registry record, the brief summary of the trial, MeSH, and the “date first received” (i.e. the date the record was first processed by ClinicalTrials.gov). The following data fields from ClinicalTrials.gov have not been republished in CENTRAL: Recruitment status, Study results, Condition, Intervention, Sponsor, Gender, Age, Phase, Enrolment, Funded by, Study type, Study design, Other IDs, Start date, Completion date, Last updated, Last verified, Acronym, Primary completion date, Outcome measures.
2.1.3.3 Records from the WHO’s International Clinical Trials Registry Platform (ICTRP)
The World Health Organization’s International Clinical Trials Registry Platform (ICTRP) (http://apps.who.int/trialsearch/Default.aspx) is a meta-register containing trials data from 17 national and international registries. Since July 2018, eligible trial registry records from ICTRP are being identified and systematically added to CENTRAL through Cochrane’s Centralized Search Service (CSS) project. As with ClinicalTrials.gov, only ICTRP records for RCTs or quasi-RCTs are being added to CENTRAL; other study designs are not included.
Process description
Backlog
The backlog (approximately 200,000 records, not including ICTRP records from ClinicalTrials.gov) was first de-duplicated against CENTRAL. Within the remaining records, those from the EU Clinical Trials Register (EU-CTR) were then de-duplicated against each other. This was because the same multicentre trial could be registered multiple times – once for each country which recruited participants for that trial. In these cases, we kept the first record created for that multicentre trial.
We then created a ‘direct feed’ search for records that were extremely likely to be describing a randomized trial. We ran the query: {(randomised OR randomized) NOT (randomised: no OR randomized: no)} in the study design and study type fields, and those with (randomised OR randomized) – see footnote to Table 2.1.a Sources searched as part of the Cochrane Centralized Search Service (CSS). This query identified 136,000 records. We manually checked around 2000 of these records to be sure that over 99% were reports of RCTs or quasi-RCTs. We sent the remaining records (around 50,000) to Cochrane Crowd for manual screening. Of these, just over 7000 RCTs were identified for CENTRAL. The backlog was cleared by December 2018.
Prospective workflow
The prospective workflow is the same as the workflow described above for the backlog. Newly identified, eligible ICTRP records are added to CENTRAL on a monthly basis and published in a new issue of CENTRAL at the end of the month.
Field mappings
Not all fields for ICTRP records are included in CENTRAL. The fields that are included are Public and Scientific titles, the URL for the registry record on ICTRP, the Key inclusion and exclusion criteria (which will be mapped to the abstract field), the date of registration (mapped to the year field), and the Study ID and the Source register.
2.1.3.4 Records from KoreaMed
KoreaMed (https://www.koreamed.org) is a database provided by the Korean Association of Medical Journal Editors that contains citations to articles published in Korean medical, dental, nursing and nutrition related journals. This database is now routinely searched and records systematically added to CENTRAL through Cochrane’s Centralized Search Service (CSS) project.
Process description
Inception to December 2013
A project led by Cochrane Australia, in partnership with KoreaMed, sought to identify all unique reports of randomized trials across all dates within the database. As part of this work a search strategy was developed and run in KoreaMed. The search strategy was:
placebo*[ALL] OR randomi*[ALL] OR randomly[ALL] OR trial*[ALL] OR ((singl* OR doubl* OR tripl* OR trebl*) AND (blind OR mask)) OR “randomized controlled trial”[PT] OR “clinical trial”[PT] OR “double blind method”[MH] OR “single blind method”[MH]
That work identified approximately 3300 unique reports of randomized trials, which were published in CENTRAL in April 2015.
January 2014 to July 2017
Between January 2014 and up to and including June 2017, all records that were added to KoreaMed within that time frame were manually screened by the Centralized Search Service team, with 1100 records submitted to CENTRAL during this time.
August 2017 onwards
From August 2017, a new process has been implemented. All KoreaMed records go through the Cochrane’s RCT machine classifier and Cochrane Crowd (crowd.cochrane.org). Records that receive a likelihood score (as described above for ClinicalTrials.gov records) of 10% or less are automatically rejected; records that receive a score of 11% or above are sent to Cochrane Crowd for manual screening.
To identify records from KoreaMed within CENTRAL, use the All Text field and the search term: HS-KOREAMED.
2.1.3.5 Records from CINAHL Plus
In November 2018 a memorandum of understanding was signed between Cochrane, Wiley and CINAHL Plus provider EBSCO (https://www.ebscohost.com/nursing/products/cinahl-databases/the-cinahl-database) to enable publication of unique CINAHL Plus records in CENTRAL. Work has begun to create a publication workflow and it is anticipated this will go live towards the end of 2019.
2.1.4 What is in the Cochrane Central Register of Controlled Trials (CENTRAL) from Specialized Registers of Cochrane Review Groups and Fields?
Most CRGs develop and maintain a Specialized Register, which aims to contain all relevant studies in their area of interest. These individual registers, together with other relevant records from other sources, are stored together as a single Cochrane Register of Studies (CRS), public records of which can be accessed by any Cochrane member logged into their Cochrane Account via the Cochrane Register of Studies Online (CRSO) (https://crso.cochrane.org/). (Note: this web address can only be accessed when logged in as above.) These public records are also published in CENTRAL in the Cochrane Library. The purpose of the Specialized Register is to assemble a repository of reports of trials relating to the scope of a CRG, to provide a reliable pool of trials for review authors that is easily retrievable, and to share this content with users of the Cochrane Library, via CENTRAL (Littlewood et al 2017). Most CRGs manage a reference-based register, where each record represents a report of a clinical trial. Where there are multiple reports of a clinical trial, as is typical, there will be multiple records for that trial. Such registers are very similar to a bibliographic database (Wieland et al 2013). Some CRGs manage a study-based register, where the reports related to each clinical trial or study have been linked together, and identified by a study name (Shokraneh and Adams 2017). In this case, there should only be one record for each clinical trial or study, with all the reports of that clinical trial or study linked to the study record. In some of these groups, the Cochrane Information Specialist also extracts metadata about studies such as the study participants, the research problem, interventions, outcomes, and study designs (Shokraneh and Adams 2017).
Specialized Registers primarily contain reports of randomized and quasi-randomized trials, however, some CRGs add other types of reports to their register, such as controlled before-and-after studies and interrupted time series (Littlewood et al 2017). Whether or not these are added to the Specialized Register will depend on the scope of the CRG. These publication types can be published in CENTRAL. CRGs can also add other reports to their register that may be useful to review authors (such as systematic reviews or background articles), but these would not be published in CENTRAL (Falzon and Trudeau 2007).
It is mandatory, for all Cochrane reviews of interventions, to search the Cochrane Review Group’s (CRG’s) Specialized Register (internally, e.g. via the Cochrane Register of Studies, or externally via CENTRAL (MECIR C24)). The Specialized Register serves to ensure that individual review authors within the CRG have easy and reliable access to trials relevant to their review topic, normally through their Cochrane Information Specialist. Records in a CRG’s Specialized Register will often contain additional metadata and other information not included in CENTRAL, so the Cochrane Information Specialist may be able to identify additional records in their Specialized Register which could not be identified by searching the Register via CENTRAL. Conversely, the search functionality of the bibliographic or other software used to manage Specialized Registers is usually less sophisticated than the search functionality available in the Cochrane Library (for example, the ability to ‘explode’ MeSH terms to include narrower, more specific terms), so a search of CENTRAL might retrieve records from the Specialized Register that may not be easily retrievable from within the Specialized Register itself. It is therefore recommended that both CENTRAL and the Specialized Register itself are searched separately to maximize retrieval.
CRGs use the methods described in Chapter 4 and the technical supplement to identify trials for their Specialized Registers. Most CRGs also have systems in place to ensure that any additional eligible reports identified by authors for their review(s) are contributed to the CRG’s Specialized Register. By sharing these registers in CENTRAL, records identified by one CRG become accessible to all others. Many Fields also develop subject-specific Specialized Registers for inclusion in CENTRAL as described above. To identify records in CENTRAL from a specific Centre, CRG or Field, it is possible to search on a Specialized Register or Handsearch code (such as SR-STROKE for records from the Cochrane Stroke Group). A list of all the Specialized Register and Handsearch codes can be found in an Appendix in the ‘CENTRAL Creation Details’ file in the Cochrane Library entitled: Cochrane Review Group, Cochrane Field or Cochrane Centre Specialized Register and handsearch codes:
https://www.cochranelibrary.com/central/central-creation.
2.2 Searching CENTRAL, MEDLINE, Embase and the Cochrane Register of Studies: specific issues
2.2.1 Searching the Cochrane Central Register of Controlled Trials (CENTRAL): specific issues
CENTRAL, accessible via the Cochrane Library or from the Cochrane Register of Studies Online (CRSO), comprises records from a wide range of sources (see Section 2.1 and subsections). The consistency and formatting of these records therefore varies. In 2013, Cochrane ran a CENTRAL “clean-up” project. The aims of this project were to clean and harmonize as many fields as possible in existing records, and to formalize standards for Cochrane Information Specialists and / or automatically apply solutions in the CRS to help prevent inconsistencies in the future.
Additionally in 2013, Cochrane formed a working group called HarmoniSR (HarmoniSR Working Group 2015). The scope of this group was initially focused on the formatting of ClinicalTrials.gov records as citations for consistent use within Cochrane Reviews and publication within CENTRAL. The scope of the group, however, expanded during 2014 onwards to include the formatting of all main record types. Despite these ongoing efforts, legitimate differences between records remain, for example, records sourced from MEDLINE will contain Medical Subject Headings (MeSH), whilst ‘native Embase’ records identified from Embase will most likely contain Emtree terms.
As of August 2019, approximately 290,000 records in CENTRAL do not have an abstract. Optimal searches will, therefore, be those that contain both MeSH and free-text terms. The 560,000 records sourced from PubMed are also best retrieved by a combination of Medical Subject Headings (MeSH) (as the Cochrane Library has a MeSH search interface) together with free-text terms. The other records, including the 412,000 records sourced from Embase, are best retrieved using free-text searches across all fields, as there is no Emtree search interface built into the Cochrane Library. Many of the records that are not sourced from PubMed or Embase (about 576,000 in CENTRAL in August 2019) either do not have abstracts or any indexing terms. To retrieve these records it is necessary to carry out a very broad search consisting of a wide range of free-text terms, which may be considered too broad to run across the whole of CENTRAL.
It is highly desirable that authors of Cochrane reviews of interventions use specially designed and tested search filters where appropriate but filters should not be used in pre-filtered databases e.g. do not use a randomized trial filter in CENTRAL (MECIR C34) or attempt to apply a limit to ‘human’ studies. All records in CENTRAL should be reports of trials in humans even though this may not be apparent from the record itself, especially for those records with no abstract.
2.2.2 Searching MEDLINE and Embase: specific issues
Irrespective of the fact that both MEDLINE and Embase have been searched systematically for reports of trials for certain years and that these reports of trials have been included in CENTRAL, as described in Sections 2.1.1 and 2.1.2, supplementary searches of both MEDLINE and Embase are recommended (as detailed below). Any such searches, however, should be undertaken in the knowledge of what searching has already been conducted to avoid duplication of effort.
Searching MEDLINE
There can be a delay of up to one month between records being indexed as trials in MEDLINE and appearing indexed as trials in CENTRAL. This is due to the Cochrane Library monthly updating cycle for CENTRAL. As a cautious approach, therefore, the most recent two months of MEDLINE should be searched, at least for records indexed as either ‘Randomized Controlled Trial’ or ‘Controlled Clinical Trial’ in the Publication Type, to identify those records recently indexed as RCTs or CCTs in MEDLINE. For further details on the search process for MEDLINE see: https://www.cochranelibrary.com/central/central-creation.
Additionally, the most recent year to be searched under the project to identify reports of trials in MEDLINE and send them back to the US National Library of Medicine for re-tagging was 2004, so records added to MEDLINE between 2005 and 2010 inclusive should be searched using one of the Cochrane Highly Sensitive Search Strategies for identifying randomized trials in MEDLINE (see Section 3.6.1). A project is planned to identify potentially missing reports from CENTRAL from this period (2005 to 2010). The project will be designed and set up as a discrete Cochrane Crowd task. (Records added to MEDLINE from 2011 onwards will have been searched as part of the Embase project described in Section 2.1.2).
Finally, for extra sensitivity, or where the use of a randomized trial filter is not appropriate, review authors should search MEDLINE for all years using appropriate free-text and thesaurus terms relevant to their review topic without any trial filter.
The MEDLINE re-tagging project described in Section 2.1.1 assessed whether the records identified were reports of trials on the basis of the title and abstract only. Any supplementary search of MEDLINE that is followed up by accessing the full text of the articles will identify additional reports of trials, most likely through the methods sections, that were not identified through the titles or abstracts alone. It is not expected, however, that accessing the full text of all articles will be routinely undertaken. For guidance on running separate search strategies in the MEDLINE-indexed versions of MEDLINE and the versions of MEDLINE containing ‘in-process’ and other non-indexed records please refer to Section 3.6.1.
Any reports of trials identified by the review author should be submitted to the Cochrane Information Specialist who can ensure that they are added to CENTRAL. Any errors, in respect of records indexed as trials in MEDLINE that on the basis of the full article are definitely not reports of trials according to the definitions used by the National Library of Medicine (NLM) (see Box 2.b), should also be reported to the Cochrane Information Specialist, so they can be referred to the NLM and corrected.
For general information about searching, which is relevant to searching MEDLINE, see Section 3 and subsections.
Searching Embase
Since 2011, the Emtree term ‘randomized controlled trial’ has been used only to index records that are reports of trials, not also for records that are about trials (as was previously the case). This change in indexing practice has made the use of the term much more precise in identifying possibly relevant studies in Embase. Users can use ‘randomized controlled trial (topic)’ [exact Ovid syntax: "randomized controlled trial (topic)"/] to help find records about RCTs. As well as the new Cochrane Embase filter (see Section 3.6.2) other search filters for searching for trials in Embase are available on the InterTASC Information Specialists’ Sub-Group website (https://sites.google.com/a/york.ac.uk/issg-search-filters-resource/filters-to-identify-randomized-controlled-trials-and).
Additionally, for extra sensitivity, or where the use of a randomized trial ‘filter’ is not appropriate, review authors should search Embase for all years using appropriate free-text and thesaurus terms relevant to their review topic without any trial filter, as described under similar circumstances for MEDLINE above.
It should be remembered that the Embase project assesses the vast majority of records identified as reports of trials on the basis of the title and abstract only. A small subset of records that have been classified Unsure by ‘Resolver’ level screeners in Cochrane Crowd do go to full-text assessment. To date this has accounted for less than 1% of all records screened for the project. Therefore, any supplementary search of Embase that is followed up by accessing the full text of the articles is likely to identify additional reports of trials, probably through the methods sections, that were not identified through the titles or abstracts alone.
There is a delay of some weeks between records being indexed in Embase and appearing in CENTRAL. The most recent months of Embase should, therefore, be searched. For more details on the Embase records workflow, go to: https://www.cochranelibrary.com/central/central-creation. Also see Table 2.1.a.
In 2011, Elsevier began to include all MEDLINE content in Embase. Before then, there had always been a sizable but not complete overlap in content between the two sources. Currently (as at August 2019), Embase includes around 3000 journals not available in MEDLINE and around 5500 journals are indexed in Embase but are also indexed in PubMed. (https://www.elsevier.com/solutions/embase-biomedical-research/embase-coverage-and-content). A search of MEDLINE, either through PubMed or through another third-party interface, is, however, still necessary. There are records in MEDLINE which have the status: PubMed-not-MEDLINE. Records with this status are “citations that will not receive MEDLINE indexing because they are for articles in non-MEDLINE journals, or they are for articles in MEDLINE journals but the articles are out of scope, or they are from issues published prior to the date the journal was selected for indexing, or citations to articles from journals that deposit their full-text articles in PMC but have not yet been recommended for indexing in MEDLINE.” (https://www.ncbi.nlm.nih.gov/books/NBK3827/table/pubmedhelp.T.status_subsets/). In addition, a recent study found that records from MEDLINE were not always retrieved when searched through Embase due to MeSH not being available in Embase (Bramer et al 2017a). Although it is, therefore, technically possible to search across all MEDLINE records in Embase (note, not all PubMed records), it is recommended that both databases be searched separately.
As noted above, in 2009 Elsevier began indexing conference abstracts for Embase and about 2.5 million conference abstracts from about 7000 conferences (as at August 2019) are now indexed in Embase. Elsevier provides a list of conferences they index for Embase, as mentioned above: (https://www.elsevier.com/solutions/embase-biomedical-research/embase-coverage-and-content). Conference abstracts can be a rich source of RCT evidence. Within Embase, these records have been indexed using automated indexing procedures, and in most cases the index terms applied automatically are about subject topics or content rather than study type. In addition, many conference abstracts have been retrospectively added to Embase, some of which have been assigned an entry date prior to the publication date of the conference abstract itself. The Embase project has made, and continues to make, efforts to identify conference records added retrospectively. It should be noted, however, that the project may not yet have identified all relevant conference publications.
2.3 Summary points
- Cochrane review authors should seek advice from their Cochrane Information Specialist on the search process.
- Authors of non-Cochrane reviews should seek advice from their medical / healthcare librarian or information specialist, with experience of conducting searches for studies for systematic reviews.
- The key databases to be searched are the Cochrane Review Group’s Specialized Register (internally, e.g. via the Cochrane Register of Studies, or externally via CENTRAL), CENTRAL, MEDLINE and Embase (if access is available to either the review author or the CRG).
- Approximately 970,000 of the 1,550,000 records in CENTRAL are from MEDLINE or Embase, so care should be taken when searching MEDLINE and Embase to avoid unnecessary duplication of effort.
- Supplementary searches of Embase and MEDLINE should be carried out as outlined in Section 2.2.2.
- Additional studies can be identified in MEDLINE and Embase by searching across the years already searched for CENTRAL, by obtaining the full article and by reading, in particular, the methods section, however, it is not expected that accessing the full text of all articles will be routinely undertaken.
3 Designing search strategies: further considerations
This section should be read in conjunction with Chapter 4, Section 4.4.
3.1 Service providers and search interfaces
Access to MEDLINE, Embase and other general and subject-specific databases is offered by several commercial service providers, via a range of search interfaces. In addition, the US National Library of Medicine, provider of MEDLINE, and Elsevier, provider of Embase, offer access to their own versions of their databases: MEDLINE through PubMed, which is available free of charge on the internet, and Embase through Elsevier directly, which is known as Embase.com and is available on subscription only. Each interface offers certain functionalities and unique features (Bethel and Rogers 2014) but more importantly the search syntax varies across the interfaces. For example, to search for the Publication Type term ‘Randomized Controlled Trial’ in MEDLINE via different search interfaces it is necessary to enter the term as:
- PT Randomized Controlled Trial (in MEDLINE on EBSCO);
- Randomized Controlled Trial.pt. (in MEDLINE on Ovid);
- DTYPE (Randomized Controlled Trial) (in MEDLINE on ProQuest); and
- Randomized Controlled Trial[pt] (in PubMed).
Although the interfaces may offer access to the same database, running the same strategy in the same database but through different interfaces may result in different search results (Schoonbaert 1996, Younger and Boddy 2009, Boeker et al 2013b, Craven et al 2014). For example, PubMed does not support proximity operators and offers limited support for phrase searching (see Section 3.5) and when using field tags to limit the search to certain parts of the record, the tags must be added after each search term or phrase and cannot be applied to all the terms by use of parentheses (brackets).
In addition to accessing bibliographic records, many service providers offer links to full-text versions of articles on other publishers’ websites, such as the PubMed ‘LinkOut’ feature. In addition, developments in the publishing industry allow users to add the DOI number, where available, after the text ‘https://doi.org/’ to retrieve the permanent location of an article on the internet.
3.2 Controlled vocabulary and text words
MEDLINE and Embase (and many other databases) can be searched using a combination of two retrieval approaches. One is based on text words (terms occurring in the title, abstract or other relevant fields) in a record. The other is based on standardized subject terms assigned to the record by indexers (specialists who appraise the article / reference and describe it by assigning terms from a specific thesaurus or controlled vocabulary). Standardized subject terms are useful because they provide a complementary way of retrieving records that may use different text words to describe the same concept and because they can provide information beyond that which is contained in the words in the title and abstract. Therefore, each concept of a robust search strategy should consist of text words together with subject terms, if the latter are available in the respective database.
It is mandatory, for Cochrane reviews of interventions, to identify appropriate controlled vocabulary (e.g. MeSH, Emtree, including ‘exploded’ terms) (see below for definition of ‘exploded’ terms (MECIR C33). When searching for studies for a systematic review, however, the extent to which subject terms are applied to references should be viewed with caution. Authors may not describe their methods or objectives well and indexers are not always experts in the subject areas or methodological aspects of the records that they are indexing. In some cases, subject terms are applied as result of automated / machine indexing and this may not be as accurate as human indexing. In addition, the available indexing terms might not correspond to the terms the searcher wishes to use. It is, therefore, mandatory, for Cochrane reviews of interventions, to identify appropriate free-text terms (considering, for example, spelling variants, synonyms, acronyms, truncation and proximity operators (MECIR C33)). This is especially important, as the indexing process in databases takes time (ranging from a few weeks to several months until a reference is fully indexed). Therefore, very current references might not yet be indexed and will consequently not be retrieved when using controlled vocabulary alone. Consideration should be given to searching indexed records and non-indexed / in-process records separately in databases such as MEDLINE and Embase which include both indexed and non-indexed content.
The approaches for identifying text words and controlled vocabulary to combine appropriately within a search strategy are presented in the following two sections and can generally be described as being subjective. Text mining is an emerging approach to identify terms in a more objective way, based on a set of relevant records on the topic (see Section 3.2.3 on text mining for term selection). Another objective method is based on similarity calculations derived from one or several known relevant articles. In MEDLINE, having identified a key article, additional relevant articles can be located by using the ‘Find Similar’ option in Ovid or the ‘Similar articles’ option in PubMed. The value of using a complementary search approach such as this feature, which is independent of the searcher’s expertise, has been described by Sampson and colleagues (Sampson et al 2016). A PubMed tutorial on the similar articles feature is available at: https://www.nlm.nih.gov/bsd/disted/pubmedtutorial/020_190.html.
3.2.1 Identifying relevant controlled vocabulary
In order to identify as many relevant records as possible, searches should include subject terms selected from the controlled vocabulary or thesaurus (‘exploded’ where appropriate - see below for definition of ‘exploded’ terms). The controlled vocabulary search terms for MEDLINE (Medical Subject Headings, known as MeSH) and Embase (Emtree) are not identical, and neither is the approach to indexing. For example, the pharmaceutical or pharmacological aspects of an Embase record are generally indexed in greater depth than the equivalent MEDLINE record, and in recent years Elsevier has increased the number of index terms assigned to each Embase record. Searches of Embase may, therefore, retrieve additional articles that were not retrieved by a MEDLINE search, even if the records were present in both databases. The converse also applies in that MEDLINE records available in Embase are indexed differently in Embase than they were originally in MEDLINE, as the MeSH terms are replaced in Embase by Emtree terms. Thus, search strategies need to be customized for each database and should ideally be run in the original database whenever possible.
Most database interfaces offer a browsing option to show the preferred subject headings. For example, interfaces to MEDLINE will usually permit browsing the Medical Subject Headings (MeSH) so that the term definition (Scope Note) and its synonyms and related terms can be searched and then inspected for relevance. Additional controlled vocabulary terms should be identified using the search tools provided with the database, such as the ‘Permuted Index’ or ‘Map Term’ under ‘Search Tools’ in Ovid or the ‘MeSH Database’ option in PubMed. As well as searching the controlled vocabulary lists, it is also common practice to identify subject headings from known relevant records. A tool which can help displaying and comparing the subject terms assigned to MEDLINE records is the ‘Yale MeSH Analyzer’ (http://mesh.med.yale.edu/) (Hocking 2017).
Many database thesauri offer the facility to ‘explode’ subject terms to include more specific terms automatically in the search. For example, a MEDLINE search using the MeSH term BRAIN INJURIES, if exploded, will automatically search not only for the term BRAIN INJURIES but also for the more specific term SHAKEN BABY SYNDROME. As articles in MEDLINE on the subject of shaken baby syndrome should only be indexed with the more specific term SHAKEN BABY SYNDROME and not also with the more general term BRAIN INJURIES, it is important that MeSH terms are ‘exploded’ wherever appropriate, in order not to miss relevant records. It is equally important, however, that MeSH terms are not ‘exploded’ where this is inappropriate, in order not to add irrelevant records unnecessarily. The same principle applies to Emtree when searching Embase and also to several other databases. For further guidance on this topic, review authors should consult their medical / healthcare librarian or information specialist.
A second option which can be applied to subject terms, is restricting the term to ‘Major Topic’ (in Ovid this feature is called ‘focus’). When this feature is used, articles are only retrieved where the subject term has been assessed by the indexer as reflecting one of the article’s major topics. This is, therefore, a precision-maximizing feature and is not recommended in the context of searching for studies for systematic reviews, as it compromises sensitivity.
It is particularly important in MEDLINE to distinguish between Publication Type terms and other related MeSH terms. For example, a report of a randomized trial should be indexed in MEDLINE with the Publication Type term ‘Randomized Controlled Trial’ whereas an article about randomized controlled trials should be indexed with the MeSH term RANDOMIZED CONTROLLED TRIALS AS TOPIC (note the word TRIALS in the latter is plural). The same applies to other indexing terms for other trials, reviews and meta-analyses. It should be noted that this distinction was also introduced into Embase for records added from 2011 onwards. The Emtree term ‘randomized controlled trial’ is used to describe the publication type of the record, whereas the Emtree term ‘randomized controlled trial (topic)’ is used for records that discuss randomized trials, but are not original reports of randomized trials. Prior to 2011, the Emtree term ‘randomized controlled trial’ was used to index both the publication type of the record and for records that discussed randomized trials as a topic.
Review authors should assume that earlier articles are even harder to identify than recent articles. For example, abstracts are not included in MEDLINE for most articles published before 1976 and, therefore, text word searches will only apply to titles. In addition, few MEDLINE indexing terms relating to study design were available before the 1990s, so text word searches relating to study design are necessary to retrieve older records.
3.2.2 Identifying relevant text words
Relevant text words (i.e. free-text terms) can be identified by checking the terms used in the title, abstract and other relevant fields (e.g. author keywords) of a few relevant references. It is important to be aware of the fact that natural language allows concepts to be expressed in different words. It is essential, therefore, to look up synonyms for each concept describing the review topic. Medical dictionaries can be used to clarify definitions and identify synonyms. The MeSH database also offers both definitions (Scope Notes) and a listing of synonyms and related terms for each MeSH term (‘Entry terms’), which lists different terms being used for a concept. Likewise, Elsevier’s Emtree thesaurus for Embase also lists synonyms for each term. A third approach for identifying text words consists of checking search strategies from other systematic reviews on a similar topic.
3.2.3 Text mining for term selection
Text mining techniques are of increasing interest in the conduct of systematic reviews generally and have been the subject of recent helpful reviews (O'Mara-Eves et al 2015, Paynter et al 2016, Stansfield et al 2017, Kohl et al 2018). Text mining encompasses a range of statistical approaches to textual analysis including simple frequency analysis of words and phrases within records, visual presentations of the inter-relationships between concepts in a literature (corpus) and the development of complex interrogation rules to identify relevant records from a corpus of records (O'Mara-Eves et al 2015, Paynter et al 2016, EUnetHTA 2017). The value of text mining can lie in its ability to process large volumes of records objectively, to assist with concept identification and to interrogate large numbers of records from many databases using a single search process. This section suggests some search-specific aspects of text mining techniques which can be combined with traditional searching approaches and also offers advice on free software.
Text mining software can be used to identify potential keywords, phrases and subject terms from within a set of relevant records. Various software packages are listed in the Systematic Review Toolbox (http://systematicreviewtools.com/).
Tools such as PubMed PubReMiner analyse the results of searches conducted in PubMed and present the words within records in order of frequency. This can aid the identification of terms, synonyms and abbreviations to test out in strategies. For databases other than MEDLINE (PubMed) frequency analysis software such as Voyant (https://voyant-tools.org/) will provide similar frequency analyses or bibliographic reference software such as EndNote (https://endnote.com/) can be used with any database records. In EndNote, frequency analysis can be achieved by using the Term Lists and the Subject Bibliography option (detailed guidance at https://sites.google.com/a/york.ac.uk/training-pages/endnote-for-text-mining).
A tool to assist with identifying relevant MeSH headings is available at the MeSH on Demand website (https://www.nlm.nih.gov/mesh/MeSHonDemand.html): it is possible, for example, to paste in a Cochrane protocol and receive suggestions of MeSH terms that relate to the topics within the text.
Tools to assist in identifying phrases and words within proximity to each other are also available in Voyant, Termine (http://www.nactem.ac.uk/software/termine/) and many other packages.
Procedures to develop search strategies routinely using text mining approaches are available (Hausner et al 2012, Hausner et al 2015, EUnetHTA 2017).
Text mining has also been used to develop methodological search filters, including the Cochrane Highly Sensitive Search Strategies for MEDLINE and Embase (Glanville et al 2006, Glanville et al 2019b) and a filter to identify overviews of systematic reviews in MEDLINE (Lunny et al 2015). Researchers are also exploring machine learning approaches to converting searches in one database to search in very different databases, such as converting PubMed searches to interrogate records in ClinicalTrials.gov (Lanera et al 2018).
Text mining may be particularly helpful when developing strategies for complex topics. Software such as VOSviewer (https://www.vosviewer.com/) can accept large numbers of records, analyse the co-occurrence of terms within records and show relationships between themes in a body of records visually. This can help with identifying, grouping and combining concepts when building strategies for complex topics (Balan et al 2014, EUnetHTA 2017).
More sophisticated text mining software which permits the development of rules for interrogating large sets of records offers opportunities for information specialists and other interested researchers to create searches across large databases containing results from many different databases and can also make use of the semantic relationships within texts to offer more precise searching. The challenges of using more sophisticated techniques include the need to acquire a working knowledge of rule building, parts of speech, ontologies and algorithms. GATE (https://gate.ac.uk/) open-source software is one example of more sophisticated text mining software which allows searchers to break down text and build new rules, to explore relationships within texts. Learning to use the software efficiently and effectively requires some investment in training and the acquisition of experience.
Text-mining tools have great potential but there are many variants and options to choose from and little guidance about what works best and when and for which questions. There is a need for more case studies and for more parallel research to show where benefits may lie. Text mining carries with it challenges in terms of documentation of the processes used and there is little guidance available on how best to report the use of text mining for strategy development.
3.3 Synonyms, related terms, variant spellings, truncation and wildcards
In order to be as comprehensive as possible, it is necessary to include a wide range of free-text terms for each of the concepts selected. This might include the use of truncation and wildcards.
It is mandatory, for Cochrane reviews of interventions, to identify appropriate spelling variants, synonyms, acronyms and truncation (MECIR C33). For example:
- synonyms: ‘pressure sore’ OR ‘decubitus ulcer’;
- related terms: ‘brain’ OR ‘head’; and
- variant spellings: ‘tumour’ OR ‘tumor’.
Database interfaces offer facilities to capture these variations through truncation and wildcards. For example:
- truncation: random* (for random or randomised or randomized or randomly etc); and
- wildcard: wom?n (for woman or women).
These features vary across different database interfaces, especially with respect to truncation length (e.g. number of characters) and position (e.g. mid-word or end-of-word), and should be checked carefully before adapting a search strategy to a different database and / or interface from that for which it was originally designed. For further details refer to the respective database help files. It should also be noted that many service providers incorporate fuzzy logic searching into their search interfaces and this automatically includes variant endings by default including singular and plural variants.
3.4 Boolean operators (AND, OR and NOT)
Boolean operators are used to join together the search terms within a search strategy. The most widely used Boolean operators are:
- AND: combines different concepts to make a set of results that is usually smaller than the smallest concept (i.e. terms from all concepts need to be present in records for them to be retrieved);
- OR: gathers terms within a concept and this usually makes the set of results larger (i.e. at least one term needs to be present in records for them to be retrieved); and
- NOT: excludes terms or concepts (one term or concept can be excluded from the set of results and the set will usually reduce in size – but see caveats below).
Generally speaking, a search strategy should build up the controlled vocabulary terms, text words, synonyms and related terms for each concept (such as the intervention), one concept at a time. Terms within a concept should normally be combined with the Boolean ‘OR’ operator: see demonstration search strategy in Box 3.h. This means records will be retrieved that contain at least one of these search terms. Sets of terms should usually be developed for the different concepts being searched such as the healthcare condition, intervention(s) and / or study design. These three concepts (sets of terms) can then be combined using the ‘AND’ operator. This combination step results in a set of records that are likely to be of the appropriate study design as well as addressing both the health condition of interest and the intervention(s) to be evaluated (see Figure 3.a). It is mandatory, for Cochrane reviews of interventions, to ensure correct use of the ‘AND’ and ‘OR’ operators (MECIR C32).
A note of caution about this approach is warranted. If a record does not contain at least one term from each of the three sets, it will not be identified. For example, if an index term has not been added to the record for the intervention and the intervention is not mentioned in the title or abstract, the record would be missed by the strategy. The best approach is to begin with as few concepts as possible and only add additional concepts if record numbers are unmanageable. So a search might begin with only one or two concepts, and the study design concept might only be added if essential.
The ‘NOT’ operator should be avoided where possible to avoid inadvertently removing from the search set any records that might be relevant. For example, when searching for records indexed as female, the use of ‘NOT male’ would remove any record that was about both males and females. NOT can be used in some situations where care is taken to ensure that relevant records are not lost, for example in the animal exclusion algorithm used within the MEDLINE search filters to identify RCTs (see Section 3.6 and subsections).
Searches to identify studies for Cochrane Reviews can sometimes be extremely long, often including over 100 search lines. It can be tedious to type in the combinations of these search sets, for example as ‘#1 OR #2 OR #3 OR #4 …. OR #100’. Some service providers offer alternatives to this. For example, in CENTRAL and Ovid it is possible to combine sets using the syntax (Littlewood et al -#100) and ‘or/1-100’ respectively. For those service providers where this is not possible, it has been recommended that the search string above could be typed in full and saved, for example, as a Word document and the requisite number of combinations copied and pasted into the search as required. Having typed the string with the # symbols as above, a second string can be generated by globally replacing the # symbol with nothing to create the string ‘1 OR 2 OR 3 OR 4 …. OR 100’ to be used for those service providers where the search interface does not use the # symbol.
Figure 3.a Combining concepts as search sets
3.5 Proximity operators (NEAR, NEXT and ADJ)
Proximity operators identify search terms which are near to each other but not necessarily directly adjacent to each other. Where the operator dictates that the search terms must be directly adjacent to each other, they are often referred to as adjacency operators. It is mandatory, for Cochrane reviews of interventions, to ensure that proximity operators are used appropriately (MECIR C33). Use of proximity operators helps to ensure that searches are more sensitive than would be the case with direct adjacency or phrase searching, and can also facilitate ease of searching where there are multiple possible variations of a phrase which would otherwise need to be typed in full.
PubMed does not support the use of proximity operators. When combining terms that appear in a phrase, the ‘AND’ Boolean operator should be considered rather than phrase searching in quotation marks in order to ensure that searches are appropriately sensitive. PubMed does, however, index lists of commonly used medical and healthcare phrases which appear in the searchable fields of PubMed records. To access a list of phrases, enter a search term in the Advanced Search Builder then click the ‘Show index list’ command next to the search box. This will bring up a list of searchable phrases, which include the specified search term. For further details, see:
https://www.ncbi.nlm.nih.gov/books/NBK3827/#pubmedhelp.Searching_for_a_phrase.
The following proximity and adjacency operators are illustrated with reference to the Cochrane Library.
NEXT
The Cochrane Library uses the proximity operator ‘NEXT’ to identify search terms which are directly adjacent to each other and in the specified order. For example, diabetes NEXT screening retrieves ‘diabetes screening’, but not ‘screening diabetes’.
‘NEXT’ functions in the Cochrane Library in the same way as searching for phrases within quotation marks such as “diabetes screening”.
NEAR
The Cochrane Library uses the operator ‘NEAR/n’ to search for search terms within a specified number of words, where n specifies the maximum number of words either search term is from the other search term in any order. For example,
- diabetes NEAR/1 screening retrieves ‘diabetes screening’ and ‘screening diabetes’
- diabetes NEAR/2 screening retrieves ‘diabetes x screening’ and ‘screening x diabetes’ where x is an intervening word
- diabetes NEAR/3 screening retrieves ‘diabetes x x screening’ and ‘screening x x diabetes’ where x is an intervening word
If the n in NEAR/n is not specified, it defaults to 6 in the Cochrane Library. Thus ‘diabetes NEAR screening’ retrieves ‘diabetes x x x x x screening’ and ‘screening x x x x x diabetes.
Syntax variation between databases
Other database interfaces use different operators, for example, ‘Nn’ in the EBSCO interface or ‘ADJn’ in the Ovid interface. Links to help pages on proximity operators for each of the main database providers are detailed at the end of this section.
It is important to note that interfaces also vary in how the number n relates to the specified search terms. In the Cochrane Library, Ovid and Embase.com interfaces n specifies the maximum number of words that either search term is from the other search term, i.e. to find a maximum of x words between two search terms n should equal x + 1. In the EBSCO, ProQuest, Scopus and Web of Science interfaces n specifies the maximum number of words between the specified search terms, i.e. to find a maximum of x words in between two search terms n should equal x. For example, if n is set to 2 it functions as shown below in the Ovid and EBSCO interfaces, respectively, where x is an intervening word:
- diabetes N2 screening retrieves ‘diabetes x x screening’ and ‘screening x x diabetes’ (EBSCO)
- diabetes ADJ2 screening retrieves ‘diabetes x screening’ and ‘screening x diabetes’ (Ovid)
If n is set to 1 in the Ovid interface it functions as shown below:
- diabetes ADJ1 screening retrieves ‘diabetes screening’ and ‘screening diabetes’
Searching using ADJ in the Ovid interface without specifying n operates in the same way as NEXT in the Cochrane Library, i.e. the search terms are retrieved but only in the specified order.
When searching using two or more search terms without quotation marks in EBSCO databases, the search terms are automatically combined using the proximity setting N5. This can be overridden by placing the terms in quotation marks, using a different proximity operator value, or combining the search terms using a Boolean operator.
Retaining the order of search terms
As noted above, the NEAR operator in the Cochrane Library and the equivalent operators used in other interfaces identify the specified search terms in any order. There is no option in the Cochrane Library for specifying the maximum number of words between search terms and retaining the specified order of the search terms. Some database providers do offer this option. For example, the EBSCO and ProQuest interfaces retain the specified order of search terms when using the ‘Wn’ and ‘pre/n’ operators, respectively, as shown below:
- diabetes W2 screening retrieves ‘diabetes x x screening’ where x is an intervening word (EBSCO)
- diabetes pre/2 screening retrieves ‘diabetes x x screening’ where x is an intervening word (ProQuest)
Help pages for proximity operators
Listed below are help links on how to use proximity operators produced by the main database providers. Some of these links go directly to the proximity operators help section and others require searching for the proximity operators section within them.
The Cochrane Library databases
https://www.wiley.com/network/cochranelibrarytraining/user-guide
EBSCO databases
https://connect.ebsco.com/s/article/How-do-I-create-a-proximity-search?language=en_US
Ovid databases
https://resourcecenter.ovid.com/site/help/documentation/ospa/en/Content/syntax.htm
ProQuest databases
https://parlipapers.proquest.com/help/parlipapers/Search_Tips.html
PubMed database (Automatic Term Mapping)
https://www.nlm.nih.gov/bsd/disted/pubmedtutorial/020_040.html
PubMed database (Searching for a Phrase)
https://www.ncbi.nlm.nih.gov/books/NBK3827/#pubmedhelp.Searching_for_a_phrase
Scopus database (Elsevier)
https://blog.scopus.com/tips-and-tricks
Web of Science databases (Clarivate Analytics)
http://images.webofknowledge.com/WOKRS58B4/help/WOS/hs_search_operators.html#dsy862-TRS_proximity
3.6 Search filters
This section should be read in conjunction with Chapter 4, Section 4.4.7.
3.6.1 The Cochrane Highly Sensitive Search Strategies for identifying randomized trials in MEDLINE
The first Cochrane Highly Sensitive Search Strategy for identifying randomized trials in MEDLINE was designed by Carol Lefebvre and published in 1994 (Dickersin et al 1994). This strategy was thereafter published in subsequent editions of this Handbook and has been adapted and updated as necessary over time. The Cochrane Highly Sensitive Search Strategies for MEDLINE, in subsequent sections, are adapted from strategies first published in 2006 as a result of a frequency analysis of MeSH terms and free-text terms occurring in the titles and abstracts of MEDLINE-indexed records of reports of randomized trials (Glanville et al 2006), using methods of search strategy design first developed by the authors to identify systematic reviews in MEDLINE (White et al 2001).
Two strategies are offered: a sensitivity-maximizing version and a sensitivity- and precision-maximizing version. It is recommended that searches for trials for inclusion in Cochrane Reviews begin with the sensitivity-maximizing version in combination with a highly sensitive subject search. If this retrieves an unmanageable number of references the sensitivity- and precision-maximizing version should be used instead. See Sections 2.1.1 and 2.2.2 for details as to how these search strategies and others have been run centrally in Cochrane over the years and relevant records included in CENTRAL, to avoid unnecessary duplication of effort.
The strategies have been updated, after re-analysis of the data used to derive those strategies, to reflect changes in search syntax and changes in indexing policy introduced by the US National Library of Medicine since the original analysis. These changes include:
- the change of the MeSH term CLINICAL TRIALS to CLINICAL TRIALS AS TOPIC; and
- no longer assigning ‘Clinical Trial’ as a Publication Type to all records indexed with ‘Randomized Controlled Trial’ or ‘Controlled Clinical Trial’ as a Publication Type.
The strategies are given in Box 3.a and Box 3.b for PubMed and in Box 3.c and Box 3.d for Ovid.
The strategies below are based on data derived from MEDLINE-indexed records and were designed to be run in MEDLINE. These strategies were not specifically designed to retrieve non-MEDLINE records in PubMed or those records in the Ovid segments: ‘in process’, other records not indexed with MeSH, and Epub Ahead of Print. It is, therefore, recommended that these strategies are run in the ‘Ovid MEDLINE(R) ALL 1946 to Month X Day X, 20XX’ Ovid segment and that the status field (ST) limit be used to isolate the MEDLINE-indexed and the non-indexed records as follows:
- all records in the database: docz.dz.
- MEDLINE status: medline.st. (i.e. MEDLINE-indexed)
- Publisher - ahead of print status: publisher.st.
- In-process & non-indexed citations: ("in data review" or in process or "pubmed not medline").st.
- Pmcbooks: nb$.bk.
The use of the various status limits and how they add up to all records in the entire MEDLINE on Ovid database (generated by the search term docz.dz.) is shown below:
Ovid MEDLINE(R) ALL <1946 to October 07, 2019>
|
# |
Searches |
Results |
|
1 |
docz.dz. |
30206097 |
|
2 |
limit 1 to medline |
26211797 |
|
3 |
limit 1 to publisher |
376666 |
|
4 |
limit 1 to ("in data review" or in process or "pubmed not medline") |
3597443 |
|
5 |
nb$.bk. |
20191 |
|
6 |
2 or 3 or 4 or 5 |
30206097 |
For identifying non-indexed records a range of truncated free-text terms would be required, such as random, placebo, trial, etc, and the search must not be limited to humans (as the records may not yet be indexed as humans).
As discussed in Section 2.1.1, MEDLINE has been searched from 1966 to 2004 inclusive, using previous versions of the Cochrane Highly Sensitive Search Strategy for identifying randomized trials, and more recent MEDLINE records (from 2011) have been searched as part of the current Embase project. All reports of trials identified in these ways (predominantly on the basis of the titles and abstracts only) are now included in CENTRAL (see Sections 2.1.1 and 2.1.2). For further guidance as to the appropriate use of these Highly Sensitive Search Strategies see Section 2.2.2.
Box 3.a Cochrane Highly Sensitive Search Strategy for identifying randomized trials in MEDLINE: sensitivity-maximizing version (2008 revision); PubMed format
|
#1 #2 #3 #4 #5 #6 #7 #8 #9 #10 #11 |
randomized controlled trial [pt] controlled clinical trial [pt] randomized [tiab] placebo [tiab] drug therapy [sh] randomly [tiab] trial [tiab] groups [tiab] #1 OR #2 OR #3 OR #4 OR #5 OR #6 OR #7 OR #8 animals [mh] NOT humans [mh] #9 NOT #10 |
PubMed search syntax (for Box 3.a above and Box 3.b below):
[pt] denotes a Publication Type term;
[tiab] denotes a word in the title or abstract;
[sh] denotes a subheading;
[mh] denotes a Medical Subject Heading (MeSH) term ‘exploded’;
[mesh: noexp] denotes a Medical Subject Heading (MeSH) term not ‘exploded’;
[ti] denotes a word in the title.
Box 3.b Cochrane Highly Sensitive Search Strategy for identifying randomized trials in MEDLINE: sensitivity- and precision-maximizing version (2008 revision); PubMed format
|
#1 #2 #3 #4 #5 #6 #7 #8 #9 #10 |
randomized controlled trial [pt] controlled clinical trial [pt] randomized [tiab] placebo [tiab] clinical trials as topic [mesh: noexp] randomly [tiab] trial [ti] #1 OR #2 OR #3 OR #4 OR #5 OR #6 OR #7 animals [mh] NOT humans [mh] #8 NOT #9 |
The search syntax is explained above under Box 3.a above.
Box 3.c Cochrane Highly Sensitive Search Strategy for identifying randomized trials in MEDLINE: sensitivity-maximizing version (2008 revision); Ovid format
|
1 2 3 4 5 6 7 8 9 10 11 |
randomized controlled trial.pt. controlled clinical trial.pt. randomized.ab. placebo.ab. drug therapy.fs. randomly.ab. trial.ab. groups.ab. 1 or 2 or 3 or 4 or 5 or 6 or 7 or 8 exp animals/ not humans.sh. 9 not 10 |
Ovid search syntax (for Box 3.c above and Box 3.d below):
.pt. denotes a Publication Type term;
.ab. denotes a word in the abstract;
.fs. denotes a ‘floating’ subheading, that is a subheading irrespective of the MeSH term to which it is attached;
exp denotes a Medical Subject Heading (MeSH) term ‘exploded’;
.sh. denotes a Medical Subject Heading (MeSH) term not ‘exploded’;
.ti. denotes a word in the title.
Box 3.d Cochrane Highly Sensitive Search Strategy for identifying randomized trials in MEDLINE: sensitivity- and precision-maximizing version (2008 revision); Ovid format
|
1 2 3 4 5 6 7 8 9 10 |
randomized controlled trial.pt. controlled clinical trial.pt. randomized.ab. placebo.ab. clinical trials as topic.sh. randomly.ab. trial.ti. 1 or 2 or 3 or 4 or 5 or 6 or 7 exp animals/ not humans.sh. 8 not 9 |
The search syntax is explained above under Box 3.c above.
3.6.2 Search filters for identifying randomized trials in Embase
As discussed in Section 2.1.2, Embase has been searched with various filters from 1980 to date (and from 1974 to 1979 for some search terms), and records of reports of trials (predominantly on the basis of screening of the titles and abstracts only) have been included in CENTRAL. Cochrane has recently funded the development of a highly sensitive search strategy for identifying reports of controlled trials in Embase (Glanville et al 2019b). This search filter was designed for the Embase database via the Ovid interface and was developed, tested and validated in 2016 (http://www.cochranelibrary.com/help/central-creation-details.html).
After the development of the filter, the Cochrane Centralized Search Service (CSS) decided to move to conducting regular searches for reports of RCTs and CCTs using the Embase.com interface, maintained by Elsevier. This move required a translation of the Ovid Embase RCT filter (Glanville et al 2019b). A proposed filter is shown in Box 3.e. Variations of this filter have been used over time to identify reports of controlled trials in Embase for inclusion in CENTRAL. Alternatively, other search filters can be identified from the ISSG search filter resource (https://sites.google.com/a/york.ac.uk/issg-search-filters-resource/filters-to-identify-randomized-controlled-trials-and).
Box 3.e Cochrane Highly Sensitive Search Strategy for identifying controlled trials in Embase: (2018 revision); Ovid format (Glanville et al 2019b)
|
1. Randomized controlled trial/ 2. Controlled clinical study/ 3. random$.ti,ab. 4. randomization/ 5. intermethod comparison/ 6. placebo.ti,ab. 7. (compare or compared or comparison).ti. 8. ((evaluated or evaluate or evaluating or assessed or assess) and (compare or compared or comparing or comparison)).ab. 9. (open adj label).ti,ab. 10. ((double or single or doubly or singly) adj (blind or blinded or blindly)).ti,ab. 11. double blind procedure/ 12. parallel group$1.ti,ab. 13. (crossover or cross over).ti,ab. 14. ((assign$ or match or matched or allocation) adj5 (alternate or group$1 or intervention$1 or patient$1 or subject$1 or participant$1)).ti,ab. 15. (assigned or allocated).ti,ab. 16. (controlled adj7 (study or design or trial)).ti,ab. 17. (volunteer or volunteers).ti,ab. 18. human experiment/ 19. trial.ti. 20. or/1-19 21. random$ adj sampl$ adj7 (“cross section$” or questionnaire$1 or survey$ or database$1)).ti,ab. not (comparative study/ or controlled study/ or randomi?ed controlled.ti,ab. or randomly assigned.ti,ab.) 22. Cross-sectional study/ not (randomized controlled trial/ or controlled clinical study/ or controlled study/ or randomi?ed controlled.ti,ab. or control group$1.ti,ab.) 23. (((case adj control$) and random$) not randomi?ed controlled).ti,ab. 24. (Systematic review not (trial or study)).ti. 25. (nonrandom$ not random$).ti,ab. 26. “Random field$”.ti,ab. 27. (random cluster adj3 sampl$).ti,ab. 28. (review.ab. and review.pt.) not trial.ti. 29. “we searched”.ab. and (review.ti. or review.pt.) 30. “update review”.ab. 31. (databases adj4 searched).ab. 32. (rat or rats or mouse or mice or swine or porcine or murine or sheep or lambs or pigs or piglets or rabbit or rabbits or cat or cats or dog or dogs or cattle or bovine or monkey or monkeys or trout or marmoset$1).ti. and animal experiment/ 33. Animal experiment/ not (human experiment/ or human/) 34. or/21-33 35. 20 not 34 |
3.6.3 Search filters for identifying randomized trials in CINAHL Plus
A search filter for identifying randomized trials in CINAHL Plus has been prepared by the Cochrane Centralized Search Service (CSS) and was published in February 2019 (Glanville et al 2019c).
Box 3.f Cochrane CINAHL Plus filter
|
S1 MH randomized controlled trials S2 MH double‐blind studies S3 MH single‐blind studies S4 MH random assignment S5 MH pretest‐posttest design S6 MH cluster sample S7 TI (randomised OR randomized) S8 AB (random*) S9 TI (trial) S10 MH (sample size) AND AB (assigned OR allocated OR control) S11 MH (placebos) S12 PT (randomized controlled trial) S13 AB (control W5 group) S14 MH (crossover design) OR MH (comparative studies) S15 AB (cluster W3 RCT) S16 MH animals+ S17 MH (animal studies) S18 TI (animal model*) S19 S16 OR S17 OR S18 S20 MH (human) S21 S19 NOT S20 S22 S1 OR S2 OR S3 OR S4 OR S5 OR S6 OR S7 OR S8 OR S9 OR S10 OR S11 OR S12 OR S13 OR S14 OR S15 S23 S22 NOT S21 |
Key
MH CINAHL Plus subject heading
+ explode subject heading
AB Word in abstract
TI Word in title
MODEL* Truncated word
W3 Within three words
3.7 Demonstration search strategies
Box 3.g provides a demonstration search strategy for CENTRAL for the topic ‘treating breast cancer with tamoxifen’. Note that it includes topic terms only and there is no limiting to humans only (a randomized trial filter is not appropriate for CENTRAL; nor is limiting to humans as CENTRAL contains only reports of trials in humans). The strategy is provided for illustrative purposes only: searches of CENTRAL for studies to include in a systematic review would have many more search terms for each of the concepts.
Box 3.h provides a demonstration search strategy for MEDLINE (Ovid format) for the topic ‘treating breast cancer with tamoxifen’. Note that both topic terms and a randomized trial filter are used for MEDLINE. The search is limited to humans. The strategy is provided for illustrative purposes only: searches of MEDLINE for studies to include in a systematic review would have many more search terms for each of the concepts.
Box 3.g Demonstration search strategy for CENTRAL, for the topic ‘treating breast cancer with tamoxifen’
|
#1 [mh “Breast Neoplasms”] #2 (breast near cancer*):ti,ab,kw #3 (breast near neoplasm*):ti,ab,kw #4 (breast near carcinoma*):ti,ab,kw #5 (breast near tumour*):ti,ab,kw #6 (breast near tumor*):ti,ab,kw #7 (Littlewood et al -#6) #8 [mh Tamoxifen] #9 tamoxifen:ti,ab,kw #10 #8 or #9 #11 #7 and #10 |
The ‘near’ operator defaults to within six words;
‘*’ indicates truncation.
Box 3.h Demonstration search strategy for MEDLINE (Ovid format), for the topic ‘treating breast cancer with tamoxifen’
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 |
randomized controlled trial.pt. controlled clinical trial.pt. randomized.ab. placebo.ab. drug therapy.fs. randomly.ab. trial.ab. groups.ab. or 1-8 exp animals/ not humans/ 9 not 10 exp Breast Neoplasms/ (breast adj6 cancer$).mp. (breast adj6 neoplasm$).mp. (breast adj6 carcinoma$).mp. (breast adj6 tumour$).mp. (breast adj6 tumor$).mp. or 12-17 exp Tamoxifen/ tamoxifen.mp. 19 or 20 11 and 18 and 21 |
The ‘adj6’ operator indicates within six words;
‘$’ indicates truncation.
As noted in the Ovid MEDLINE 2019 Database Guide, under ‘Default Fields for Unqualified Searches (MP)’: searching for a term without specifying a field in Advanced search, or specifying .mp., defaults to the following ‘multi-purpose’ (.mp.) fields for this database: ti,ab,ot,nm,hw,fx,kf,ox,px,rx,ui,sy.
The above field labels stand for: Title (TI), Abstract (AB), Original Title (OT), Name of Substance Word (NM), Subject Heading Word (HW), Floating Sub-Heading Word (FX), Keyword Heading Word (KF), Organism Supplementary Concept Word (OX), Protocol Supplementary Concept Word (PX), Rare Disease Supplementary Concept Word (RX), Unique Identifier (UI), Synonyms (SY).
http://ospguides.ovid.com/OSPguides/medline.htm.
3.8 Adapting search strategies across databases / sources and interfaces
Search strategies need to be customized for each database and search interface. Special caution is warranted when adapting a search strategy developed for a specific database in a specific interface to other databases and / or interfaces. This process requires a thorough knowledge of the specifications of both the new database and the new interface, including the controlled vocabulary being used to index the database’s content and the availability of Boolean and proximity operators, as well as the specific syntax for wildcards and truncation and definitions of date fields. These vary across databases and interfaces and need to be taken into account before running a strategy. Searchers should be particularly vigilant with respect to wildcard and truncation symbols, which in some cases have the opposite meaning in different database interfaces. Additionally, a search for health economics in a general healthcare database such as MEDLINE will require different natural language (free-text) terminology / search terms from the terminology required in a specialized economics database. Review authors are, therefore, encouraged to work together with their healthcare librarian or Cochrane Information Specialist, who can provide advice on the accuracy of adaptations carried out by the review authors themselves or may be able to provide adaptations of the principal, generally MEDLINE, search strategy into the databases and trials registers, which will be searched for the review. Some attempts have been made to simplify through automation the adaptation of search syntax across service providers:
Bond University Centre for Research in Evidence-Based Practice Systematic Review Accelerator Polyglot application project http://sr-accelerator.com/#/polyglot
Erasmus University Medical Centre (Bramer et al 2017b) http://www.stationsweb.nl/emcmb_cursus/bestanden/macros.html
MEDLINE Transpose from the College of Physicians and Surgeons of British Columbia (CPSBC) and the Collaboration for Leadership in Applied Health Research and Care South West Peninsula (PenCLAHRC) https://medlinetranspose.github.io/about.html (Wanner and Baumann 2018).
None of the above, however, addresses the complexities outlined above regarding differences in natural language (free-text) terminology or controlled vocabulary.
With respect to date fields, the table below indicates the equivalent date fields between Ovid and PubMed. For example, it is important to note that the Publication Date (DP) field in PubMed (for the date that the article was published) is not equivalent to the Year of Publication (YR) field in Ovid MEDLINE – see Table 3.8.a.
Table 3.8.a Equivalent date fields between Ovid and PubMed
|
PubMed Search |
Ovid Search |
|
1950:2015[epdat] |
EP - Electronic Date of Pub.: 19500101:20151231.(ep). |
|
("1950"[Date - Publication] : "2015"[Date - Publication]) |
YR or EP: 1950:2015.(yr). or 19500101:20151231.(ep). |
|
("1950"[Date - MeSH] : "2015"[Date - MeSH]) |
DA - MeSH date: 19500101:20151231.(da). |
|
("1950"[Date - Entrez] : "2015"[Date - Entrez]) |
EZ - Entrez date: 19500101:20151231.(ez). |
|
("1950"[Date - Create] : "2015"[Date - Create]) |
DT - Create date: 19500101:20151231.(dt). |
|
("1950"[Date - Completion] : "2015"[Date - Completion]) |
ED - entry date: 19500101:20151231.(ed). |
3.9 Identifying fraudulent studies, other retracted publications, errata and comments: further considerations
This section should be read in conjunction with Chapter 4, Section 4.4.6. It is mandatory, for authors of Cochrane reviews of interventions, to examine any relevant retraction statements and errata for information (MECIR C48). Identifying retraction statements and published errata or comments (and their associated original retracted articles or corrected articles) can help to avoid errors that impact on the overall estimates in systematic reviews. It is essential at the original search stage to ascertain whether any retractions or errata have been published for studies to be included in the original review and also at the update stage to ascertain whether any retractions or errata have been published subsequently for studies previously included in the original review. There is an increasing awareness of the importance of not including retracted studies or those with significant errata in systematic reviews and how best to avoid this (Royle and Waugh 2004, Wright and McDaid 2011, Decullier et al 2014). A recent study, however, showed that even when review authors suspect research misconduct, including data falsification, in the trials that they are considering including in their systematic reviews, they do not always report it (Elia et al 2016).
Reports of studies indexed in MEDLINE that have been retracted (as fraudulent or for other reasons) will have the Publication Type term ‘Retracted Publication’ added to the record (since 1989). The article giving notice of the retraction (the retraction notice) will have the Publication Type term ‘Retraction of Publication’ assigned (since 1991).
How to search for retraction notices and retracted publications in Ovid MEDLINE:
- retracted publication.pt. or retraction of publication.pt.
How to search for retraction notices and retracted publications in PubMed:
- retracted publication [pt] OR retraction of publication [pt]
The above searches could be supplemented with a free-text search of ‘retracted’ or ‘retraction’ limited to the title, to pick up records not (yet) indexed as such but this will inevitably result in false positives, i.e. irrelevant records.
Retraction notices indexed in Embase until April 2017 were identified by the Publication Type ‘erratum’ and were additionally indexed with the Preferred Term ‘retracted article’. There was no link, prior to April 2017, back from the retraction notice to the original retracted article, as there is in MEDLINE.
How to search for retraction notices and retracted publications in Ovid Embase:
- Erratum.pt. or Retracted article/ or Tombstone.pt. or yes.nr.
As above for MEDLINE, the above search in Embase could be supplemented with a free-text search of ‘retracted’ or ‘retraction’ limited to the title, to pick up records not (yet) indexed as such but this will inevitably result in false positives, i.e. irrelevant records.
Prior to any decision being taken to retract an article, articles may be published that refer to an original article and raise concerns of this sort. A new MeSH Publication Type was introduced in 2018 to cover this: Expression of Concern. This is defined in the Scope Note as: “A notification about the integrity of a published article that is typically written by an editor and should be labelled prominently in the item title. It is the responsibility of the editor to initiate appropriate investigative procedures, discover the outcome of the investigation, and notify readers of that outcome in a subsequent published item. The outcome may require the publication of a retraction notice.”
To search for “expressions of concern” prior to 2018, search for the phrase “expression of concern”.
Search in Ovid as:
expression of concern.pt. or expression of concern.af.
Search in PubMed as:
“expression of concern”[Publication Type] OR “expression of concern”[All Fields]
As noted above, MEDLINE/PubMed, reports of randomized trials that have been retracted and indexed as such in the MEDLINE, will include the ‘Retracted Publication’ term in the Publication Type field (since 1989). This is also the case for those retracted articles in CENTRAL which are sourced from MEDLINE. This is not, however, the case for the majority of records from Embase (prior to 2017) or from other sources.
In addition, articles may have been partially retracted (previously indexed in MEDLINE as Partial Retraction but since 2016 indexed as Erratum), corrected through a published erratum or may have been corrected and re-published in full. It is therefore important to search MEDLINE for the latest version of the citations to the records for the (previously) included studies when updating a review. In some display formats of some versions of MEDLINE the retracted publication, erratum and comment statements are included in the citation data together with the title and are, therefore, highly visible. This is not, however, always the case so care should be taken to ensure that this information is always retrieved in all searches by downloading the appropriate fields together with the citation data.
Retraction Watch is a resource listing retracted publications (since late 2010). Review authors and others interested in keeping abreast of this area can subscribe to their blog by email (approximately 100,000 subscribers as at January 2018) and search their blog and archives by category (http://retractionwatch.com/).
3.10 Summary points
- Cochrane review authors should seek advice from their Cochrane Information Specialist on designing search strategies.
- Authors of non-Cochrane reviews should seek advice from their medical / healthcare librarian or information specialist, with experience of conducting searches for studies for systematic reviews.
- Avoid too many different search concepts but use a wide variety of synonyms and related terms.
- Appropriate controlled vocabulary (e.g. MeSH, Emtree, including ‘exploded’ terms) and free-text terms should be identified (considering, for example, spelling variants, synonyms, acronyms, truncation and proximity operators).
- Ensure correct use of the ‘AND’ and ‘OR’ operators.
- Avoid use of the ‘NOT’ operator in combining search sets.
- Specially designed and tested search filters should be used where appropriate including the Cochrane Highly Sensitive Search Strategies for identifying randomized trials in MEDLINE, Embase and CINAHL Plus.
- Do not use filters in pre-filtered databases e.g. do not use a randomized trial or human studies filter in CENTRAL or a systematic review filter in a database consisting solely of systematic reviews.
- For identifying randomized trials in MEDLINE, begin with a highly sensitive search filter such as the sensitivity-maximizing version of the Cochrane Highly Sensitive Search Strategy. If this retrieves an unmanageable number of references, use the sensitivity- and precision-maximizing version instead. (See Sections 2.1.1 and 2.2.2 for details as to how these search strategies have already been run centrally in Cochrane over the years and relevant records included in CENTRAL, to avoid unnecessary duplication of effort.)
- Searches designed for a specific database and service provider will need to be adapted for use in another database or service provider.
- Ensure awareness of any retracted publications (e.g. fraudulent publications), errata and comments.
- Consideration should be given to searching indexed records and non-indexed / in-process records separately in databases such as MEDLINE and Embase which include both indexed and non-indexed content.
4 Managing references
4.1 Reference Management software
Specially designed bibliographic or reference management software such as EndNote (https://endnote.com/), Mendeley (https://www.mendeley.com/), RefWorks (https://www.proquest.com/products-services/refworks.html) and Zotero (https://www.zotero.org/) is useful and relatively easy to use to keep track of references to and other records of studies (Lorenzetti and Ghali 2013). Reference management software varies in terms of cost, operating system, and ease of database and record sharing, among other characteristics. The choice of which software to use is likely to be influenced by what is available and thus supported at the review author’s institution. There are currently (March 2019) 37 different software tools listed in the reference management section of the Systematic Review Toolbox at: http://systematicreviewtools.com/. For a comparison of the main products see en.wikipedia.org/wiki/Comparison_of_reference_management_software.
Reference management software usually provides import file formats (import filters) that allow text files exported from sources such as CENTRAL, CINAHL, Embase, MEDLINE, PsycINFO, PubMed and others to be imported into the reference management database. Some reference management software can also be used to search sources such as PubMed from within the database of citations and to import retrieved records directly from those sources. Using reference management software to carry out complex searches, such as those for identifying studies for systematic reviews, is, however, discouraged (Gomis et al 2008).
Reference management software facilitates storage of information about the methods and process of a search. For example, unused record fields can be used to store information such as 1) the name of the database or other source details from which a trial record was identified, 2) when and from where a document was ordered and the date of document receipt, 3) when and with whom the search results were shared, and 4) whether the study associated with a record / document was included in or excluded from a review and, if excluded, the reasons for exclusion.
Increasingly software is being developed to manage a range of functions within the systematic review process and many of these also have some level of reference management capacity. Further information about these software tools is available from the Systematic Review Toolbox at http://systematicreviewtools.com/.
4.2 Which fields to download
In addition to the fields that are essential for identifying a reference (e.g. author, title, source, year) several additional key fields should be considered for downloading from databases where they are available. Some of these key fields are listed below. The list below is intended, where possible, to be generic across databases. For the full range of fields in PubMed, see https://www.nlm.nih.gov/bsd/mms/medlineelements.html.
Abstract: abstracts can be used to eliminate clearly irrelevant reports, obviating the need to obtain the full text of those reports or to return to the bibliographic database at a later time.
Accession number / unique identifier: it is advisable to allocate an unused field or fields to store the unique identifier(s) / accession number(s) of records downloaded, such as the PubMed ID number (PMID). This allows subsequent linkage to the full database record and also facilitates information management such as duplicate detection and removal (i.e. de-duplication).
Affiliation / address: may include the institutional affiliation and / or email address of the author / investigator.
Article identifier / digital object identifier (DOI): can be used to cite and link to the full record.
Author identifier: can be used to disambiguate authors with similar names. The identifier may be an ORCID (https://orcid.org/about/what-is-orcid/mission?), an International Standard Name Identifier (ISNI) http://www.isni.org/, or from the Virtual International Authority File (VIAF) http://viaf.org/.
Clinical trial number: if the record contains a clinical trial number, such as those assigned by the ClinicalTrials.gov or ISRCTN schemes, or a number allocated by the sponsor of the trial, these should be downloaded to aid linking of trial reports to the original studies. In PubMed, the Secondary Source ID field [SI] contains information from secondary sources such as ClinicalTrials.gov and ISRCTN. Similarly, in Ovid MEDLINE, the Secondary Source Linking (SL) field contains the URL to ClinicalTrials.gov and ISRCTN resources where these are mentioned in MEDLINE records. In Embase, the Clinical Trial Number (CN) field contains clinical trial numbers associated with the record.
Index terms / thesaurus terms / keywords: These help indicate why records were retrieved if the title and abstract lack detail.
Investigator name: this field contains personal names of individuals (e.g. collaborators and investigators) who are not authors of the article but rather are listed in the article as members of a collective / corporate group that is an author of the article.
Language: this is the language (or languages) of publication of the original document.
Location identifier: this field may also contain a Digital Object Identifier (DOI).
Original title: if the original title of the document is not in English and the original title is available, then both titles should be downloaded into separate database fields, to aid correct identification of the reference and de-duplication. See also Transliterated title below.
Other term: this field contains largely non-MeSH subject terms (also referred to as Keywords) that describe the content of the article. Author-supplied keywords are included here in PubMed (since 2013).
Registry Number / EC Number and Substance Name: these fields provide supplementary subject information regarding substances (chemicals, drugs and enzymes).
Transliterated title: in PubMed, this field contains the original title (or, where available, the transliterated title) of each record originally published in a non-English language. This field can be useful for de-duplication.
Comments, corrections, errata, retractions and updates:
It is mandatory, for Cochrane reviews of interventions, to examine any relevant retraction statements and errata for information (MECIR C48). All fields that relate to subsequently published comments, corrections, errata, retractions and updates should be selected for inclusion in the download, so that any impact of these subsequent publications can be taken into account. The MECIR standard specifies: “Care should be taken to ensure that this information is retrieved in all database searches by downloading the appropriate fields, together with the citation data”. For example, the most important fields to consider, in relation to comments, errata etc, together with their field labels in PubMed, are provided in Box 4.a.
Box 4.a Important field labels in PubMed in relation to comments, retractions etc
|
CIN: ‘Comment in’ CON: ‘Comment on’ CRI: ‘Corrected and republished in’ CRF: ‘Corrected and republished from’ EIN: ‘Erratum in’ EFR: ‘Erratum for’ ECI: Expression Of Concern In ECF: Expression Of Concern For RIN: ‘Retraction in’ ROF: ‘Retraction of’ RPI: ‘Republished in’ RPF: ‘Republished from’ UIN: ‘Update in’ UOF: ‘Update of’ |
See: https://www.nlm.nih.gov/bsd/mms/medlineelements.html
The above list is provided as an example of the relevant fields in PubMed and as an indicator of the equivalent fields in other databases and service providers.
4.3 De-duplicating references
Because searching to inform systematic reviews is intended to be extensive, thousands of records may be retrieved from multiple sources. References to the same article may be downloaded multiple times from different sources and duplicates can even be found within individual databases. The identification and elimination of duplicate records (de-duplication) reduces unnecessary work during the screening phase. Removing duplicate records from the pool of retrieved references is also necessary if the total number of records identified through database searching (in addition to the total number of additional records identified through other sources) is to be reported correctly in the PRISMA flow diagram together with the total number of records after the duplicates have been removed (Liberati et al 2009). Many Cochrane Information Specialists de-duplicate records so that review authors see only search results that have already been de-duplicated.
Formatting of citation information often varies across sources, and automated identification of duplicate references from within reference management software may lead to false positives (removing non-duplicate records) and false negatives (retaining duplicate records). Meanwhile, de-duplication through visual examination of each record is time-consuming and often impractical. Several strategies have been developed to address these issues. Methods for modifying duplicate detection algorithms within reference management software have been developed and tested (Kwon et al 2015, Bramer et al 2016b). An online method to identify search results that are duplicates of PubMed citations has been reported (Sampson et al 2006). Open-source software programs for online duplicate detection have also been developed (Jiang et al 2014, Rathbone et al 2015). There is no consensus on the optimal method for duplicate detection, and the most appropriate method will most likely depend upon the size of the combined dataset, the number and output format of the resources searched, and the skill and comfort level of the operator. A combination of automated methods and visual inspection is often used.
After de-duplication of search results, records may be screened for inclusion from within the reference management database. Alternatively, the records may be exported into dedicated screening software or into systematic review production software that includes screening capabilities. If screening is carried out within the reference management database, records for the included and excluded studies can be exported and uploaded into systematic review software such as RevMan. Instructions for importing references into RevMan can be found at: https://community.cochrane.org/help/tools-and-software/revman-5/support-revman-5/revman-5-faq.
The decision whether to screen within reference management software or within dedicated screening or review production software will most likely depend upon the number of retrieved references, access to various tools and review author preference.
4.4 Summary points
- Cochrane review authors should seek advice from their Cochrane Information Specialist on managing references.
- Authors of non-Cochrane reviews should seek advice from their medical / healthcare librarian or information specialist, with experience of managing references for systematic reviews.
- Use of reference management software is recommended.
- Ensure that all the necessary fields are downloaded.
- Remove duplicate references before screening.
- Either screen references within the reference management software and export references for the included and excluded studies into systematic review software, or export references to specialized screening software.
5 Supplement information
Authors: Carol Lefebvre, Julie Glanville, Simon Briscoe, Anne Littlewood, Chris Marshall, Maria-Inti Metzendorf, Anna Noel-Storr, Tamara Rader, Farhad Shokraneh, James Thomas and L. Susan Wieland on behalf of the Cochrane Information Retrieval Methods Group
6 Acknowledgements
This chapter has been developed from sections of previous editions of this Handbook co-authored since 1995 by Kay Dickersin, Julie Glanville, Kristen Larson, Carol Lefebvre and Eric Manheimer. Many of the sources listed in this technical supplement, the accompanying Appendix and Chapter 4 of the Cochrane Handbook for Systematic Reviews of Interventions have been brought to our attention by a variety of people over the years and we should like to acknowledge this. We should like to acknowledge: Robin Featherstone, Information Specialist, Cochrane Editorial and Methods Department, Ruth Foxlee, (formerly) Information Specialist, Cochrane Editorial Unit; Miranda Cumpston, (formerly) Head of Learning and Support, Cochrane Central Executive; Colleen Finley, Product Manager, John Wiley and Sons, for checking sections relating to searching the Cochrane Library; the (UK) National Institute for Health and Care Excellence and the German Institute for Quality and Efficiency in Health Care (IQWiG) for support in identifying some of the references; the (US) Agency for Healthcare Research and Quality (AHRQ) Effective Healthcare Program Scientific Resource Center Article Alert service; Tianjing Li, Co-Convenor, Comparing Multiple Interventions Methods Group, for text and references that formed the basis of the re-drafting of parts of the section on Selecting studies; Lesley Gillespie, Cochrane author and former Editor and Trials Search Co-ordinator of the Cochrane Bone, Joint and Muscle Trauma Group, for copy-editing an early draft; The Cochrane Information Specialist Executive, the Cochrane Information Specialists’ Support Team, Cochrane Information Specialists and members of the Cochrane Information Retrieval Methods Group for comments on drafts.
7 References
Abhijnhan A, Surcheva Z, Wright J, Adams CE. Searching a biomedical bibliographic database from Bulgaria: the ABS database. Health Information and Libraries Journal 2007; 24: 200-203.
Adam GP, Springs S, Trikalinos T, Williams JW, Jr., Eaton JL, Von Isenburg M, Gierisch JM, Wilson LM, Robinson KA, Viswanathan M, Middleton JC, Forman-Hoffman VL, Berliner E, Kaplan RM. Does information from ClinicalTrials.gov increase transparency and reduce bias? Results from a five-report case series. Systematic Reviews 2018; 7: 59.
Al-Hajeri AA, Fedorowicz Z, Amin FA, Eisinga A. The handsearching of 2 medical journals of Bahrain for reports of randomized controlled trials. Saudi Medical Journal 2006; 27: 526-530.
Almerie MQ, Matar H, Jones V. Searching the Polish Medical Bibliography (Polska Bibliografia Lekarska) for trials. Health Information and Libraries Journal 2007; 24: 283-286.
Anderson ML, Chiswell K, Peterson ED, Tasneem A, Topping J, Califf RM. Compliance with results reporting at ClinicalTrials.gov. New England Journal of Medicine 2015; 372: 1031-1039.
Atsawawaranunt K, Adams CE, Roberts S. Thai online bibliographical biomedical databases [Poster] 17th Cochrane Colloquium; 2009; Singapore.
Atsawawaranunt K, Adams CE, Roberts S. Searching for randomised controlled trials and clinical controlled trials in Thai online bibliographical biomedical databases. Health Information and Libraries Journal 2011; 28: 68-76.
Balan PF, Gerits A, Vanduffel W. A practical application of text mining to literature on cognitive rehabilitation and enhancement through neurostimulation. Frontiers in Systems Neuroscience 2014; 8: 182.
Barnabas J, Yamuna B, Parthasarathy V, Venkatesh S, Tharyan P. Access to evidence from countries in South Asia: the South Asian Database of Controlled Clinical Trials and the South Asian Cochrane Network and Centre’s Digital Library. 17th Cochrane Colloquium; 2009; Singapore. https://abstracts.cochrane.org/2009-singapore/access-evidence-countries-south-asia-south-asian-database-controlled-clinical-trials.
Beatty S. Breaking the 1996 barrier: Scopus adds nearly 4 million pre-1996 articles and more than 83 million references 2015. https://blog.scopus.com/posts/breaking-the-1996-barrier-scopus-adds-nearly-4-million-pre-1996-articles-and-more-than-83.
Becker JE, Krumholz HM, Ben-Josef G, Ross JS. Reporting of results in ClinicalTrials.gov and high-impact journals. JAMA 2014; 311: 1063-1065.
Beckles Z, Glover S, Ashe J, Stockton S, Boynton J, Lai R, Alderson P. Searching CINAHL did not add value to clinical questions posed in NICE guidelines. Journal of Clinical Epidemiology 2013; 66: 1051-1057.
Bethel A, Rogers M. A checklist to assess database-hosting platforms for designing and running searches for systematic reviews. Health Information and Libraries Journal 2014; 31: 43-53.
Blumle A, Antes G. Ten years of handsearching in Germany: results and future prospects [poster]. 13th Cochrane Colloquium; 2005; Melbourne, Australia. http://abstracts.cochrane.org/2005-melbourne/ten-years-handsearching-germany-results-and-future-prospects.
Boeker M, Vach W, Motschall E. Google Scholar as replacement for systematic literature searches: good relative recall and precision are not enough. BMC Medical Research Methodology 2013a; 13: 131.
Boeker M, Vach W, Motschall E. Time-dependent migration of citations through PubMed and OvidSP subsets: a study on a series of simultaneous PubMed and OvidSP searches. Studies in Health Technology and Informatics 2013b; 192: 1196.
Boluyt N, Tjosvold L, Lefebvre C, Klassen TP, Offringa M. The usefulness of systematic review search strategies in finding child health systematic reviews in MEDLINE. Archives of Pediatrics and Adolescent Medicine 2008; 162: 111-116.
Bonfill X, Osorio D, Posso M, Sola I, Rada G, Torres A, Garcia Dieguez M, Pina-Pozas M, Diaz-Garcia L, Tristan M, Gandarilla O, Rincon-Valenzuela DA, Marti A, Hidalgo R, Simancas-Racines D, Lopez L, Correa R, Rojas-De-Arias A, Loza C, Gianneo O, Pardo H, Iberoamerican Cochrane Network. Identification of biomedical journals in Spain and Latin America. Health Information and Libraries Journal 2015; 32: 276-286.
Booth A. Cochrane or cock-eyed? How should we conduct systematic reviews of qualitative research? Qualitative Evidence-based Practice Conference, Taking a Critical Stance; 2001; Coventry. http://www.leeds.ac.uk/educol/documents/00001724.htm.
Boynton J, Glanville J, McDaid D, Lefebvre C. Identifying systematic reviews in MEDLINE: developing an objective approach to search strategy design. Journal of Information Science 1998; 24: 137-157.
Bramer WM, Giustini D, Kramer BMR, Anderson PF. The comparative recall of Google Scholar versus PubMed in identical searches for biomedical systematic reviews: a review of searches used in systematic reviews. Systematic Reviews 2013; 2: 115.
Bramer WM. Variation in number of hits for complex searches in Google Scholar. Journal of the Medical Library Association 2016; 104: 143-145.
Bramer WM, Giustini D, Kramer BM. Comparing the coverage, recall, and precision of searches for 120 systematic reviews in Embase, MEDLINE, and Google Scholar: a prospective study. Systematic Reviews 2016a; 5: 39.
Bramer WM, Giustini D, de Jonge GB, Holland L, Bekhuis T. De-duplication of database search results for systematic reviews in EndNote. Journal of the Medical Library Association 2016b; 104: 240-243.
Bramer WM, Rethlefsen ML, Kleijnen J, Franco OH. Optimal database combinations for literature searches in systematic reviews: a prospective exploratory study. Systematic Reviews 2017a; 6: 245.
Bramer WM, Rethlefsen ML, Mast F, Kleijnen J. Evaluation of a new method for librarian-mediated literature searches for systematic reviews. Research Synthesis Methods 2017b; 9: 510-520.
Brand-de Heer DL. A comparison of the coverage of clinical medicine provided by PASCAL BIOMED and MEDLINE. Health Information and Libraries Journal 2001; 18: 110-116.
Brassey JR. Turning Research Into Practice (TRIP). Journal of the Medical Library Association 2007; 95: 215-216.
Briscoe S, Cooper C. The British Nursing Index and CINAHL: a comparison of journal title coverage and the implications for information professionals. Health Information and Libraries Journal 2014; 31: 195-203.
Briscoe S. A review of the reporting of web searching to identify studies for Cochrane systematic reviews. Research Synthesis Methods 2018; 9: 89-99.
Chibuzor M, Meremikwu M. Preliminary assessment of handsearching programmes for randomized controlled trials in Nigeria [Poster]. 17th Cochrane Colloquium; 2009; Singapore.
Chokkalingam A, Scherer R, Dickersin K. Agreement of data in abstracts compared to full publications. Contemporary Clinical Trials 1998; 19: S61-S62.
Clark OAC, Castro AA, Atallah AN. Searching LILACS database improves systematic reviews. 6th Cochrane Colloquium; 1998; Baltimore (MD), USA.
Clark OAC, Castro AA. Cochrane reviews must use LILACS database-like source of articles. 9th Annual Cochrane Colloquium Abstracts; 2001; Lyon.
Clark OAC, Castro AA. Searching the Literatura Latino Americana e do Caribe em Ciencias da Saude (LILACS) database improves systematic reviews. International Journal of Epidemiology 2002; 31: 112-114.
Cohen JF, Korevaar DA, Wang J, Spijker R, Bossuyt PM. Should we search Chinese biomedical databases when performing systematic reviews? Systematic Reviews 2015; 4: 23.
Cooper C, Lovell R, Husk K, Booth A, Garside R. Supplementary search methods were more effective and offered better value than bibliographic database searching: A case study from public health and environmental enhancement. Research Synthesis Methods 2017a; 9: 195-223.
Cooper C, Booth A, Britten N, Garside R. A comparison of results of empirical studies of supplementary search techniques and recommendations in review methodology handbooks: a methodological review. Systematic Reviews 2017b; 6: 234.
Craane B, Dijkstra PU, Stappaerts K, De Laat A. Methodological quality of a systematic review on physical therapy for temporomandibular disorders: influence of hand search and quality scales. Clinical Oral Investigations 2012; 16: 295-303.
Craven J, Jefferies J, Kendrick J, Nicholls D, Boynton J, Frankish R. A comparison of searching the Cochrane Library databases via CRD, Ovid and Wiley: implications for systematic searching and information services. Health Information and Libraries Journal 2014; 31: 54-63.
Dal-Re R, Ross JS, Marusic A. Compliance with prospective trial registration guidance remained low in high-impact journals and has implications for primary end point reporting. Journal of Clinical Epidemiology 2016; 75: 100-107.
De Oliveira GS, Jr., Jung MJ, McCarthy RJ. Discrepancies between randomized controlled trial registry entries and content of corresponding manuscripts reported in anesthesiology journals. Anesthesia and Analgesia 2015; 121: 1030-1033.
Decullier E, Huot L, Maisonneuve H. What time-lag for a retraction search on PubMed? BMC Research Notes 2014; 7: 395.
Devine J, Egger-Sider F. Going Beyond Google Again: Strategies for Using and Teaching the Invisible Web. London: Facet Publishing; 2013.
Dickersin K, Scherer R, Lefebvre C. Identifying relevant studies for systematic reviews. BMJ 1994; 309: 1286-1291.
Dickersin K, Manheimer E, Wieland S, Robinson KA, Lefebvre C, McDonald S. Development of the Cochrane Collaboration's CENTRAL Register of controlled clinical trials. Evaluation and the Health Professions 2002; 25: 38-64.
Diezel K, Pharoah FM, Adams CE. Abstracts of trials presented at the Vth World Congress of Psychiatry (Mexico, 1971): a cohort study. Psychological Medicine 1999; 29: 491-494.
Dogpile.com. Different engines, different results: Web searches not always finding what they're looking for 2007. http://www.assessmentpsychology.com/metasearch-analysis.pdf.
Doshi P. FDA to begin releasing clinical study reports in pilot programme. BMJ 2018; 360: k294.
Earley A, Lau J, Uhlig K. Haphazard reporting of deaths in clinical trials: a review of cases of ClinicalTrials.gov records and matched publications; a cross-sectional study. BMJ Open 2013; 3: e001963.
Egger M, Juni P, Bartlett C, Holenstein F, Sterne J. How important are comprehensive literature searches and the assessment of trial quality in systematic reviews? Empirical study. Health Technology Assessment 2003; 7: 1-76.
Eisinga A, Siegfried N, Clarke M. The sensitivity and precision of search terms in Phases I, II and III of the Cochrane Highly Sensitive Search Strategy for identifying reports of randomized trials in MEDLINE in a specific area of health care - HIV/AIDS prevention and treatment interventions. Health Information and Libraries Journal 2007; 24: 103-109.
Elia N, von Elm E, Chatagner A, Popping DM, Tramer MR. How do authors of systematic reviews deal with research malpractice and misconduct in original studies? A cross-sectional analysis of systematic reviews and survey of their authors. BMJ Open 2016; 6: e010442.
EUnetHTA. Process of information retrieval for systematic reviews and health technology assessments on clinical effectiveness (Version 1.2) Germany: European network for Health Technology Assessment; 2017. https://www.eunethta.eu/wp-content/uploads/2018/01/Guideline_Information_Retrieval_V1-2_2017.pdf.
Eysenbach G, Tuische J, Diepgen TL. Evaluation of the usefulness of Internet searches to identify unpublished clinical trials for systematic reviews. Medical Informatics and the Internet in Medicine 2001; 26: 203-218.
Falzon L, Trudeau KJ. Developing a database of behavioural medicine interventions. Health Information and Libraries Journal 2007; 24: 257-266.
Farace DJ, Frantzen J. Third International Conference on Grey Literature: Perspectives on the design and transfer of scientific and technical information; 1997; Luxembourg. Amsterdam: TransAtlantic; 1997.
Farace DJ, Frantzen J. Sixth International Conference on Grey Literature: Work on Grey in Progress; 2004; New York. Amsterdam: GreyNet, Grey Literature Network Service; 2005.
Farrah K, Mierzwinski-Urban M. Almost half of references in reports on new and emerging nondrug health technologies are grey literature. Journal of the Medical Library Association 2019; 107: 43-48.
Fedorowicz Z, Amin F, Eisinga A, Al-Sayyad J. Handsearching for 'buried' randomized trials in Bahrain medical journals [Poster]. 13th Cochrane Colloquium; 2005; Melbourne, Australia.
Gandhi R, Jan M, Smith HN, Mahomed NN, Bhandari M. Comparison of published orthopaedic trauma trials following registration in Clinicaltrials.gov. BMC Musculoskeletal Disorders 2011; 12: 278.
Garfield E. The evolution of the Science Citation Index. International Microbiology 2007; 10: 65-69.
Gill CJ. How often do US-based human subjects research studies register on time, and how often do they post their results? A statistical analysis of the Clinicaltrials.gov database. BMJ Open 2012; 2: e001186.
Glanville J, Lefebvre C. Identifying systematic reviews: key resources. ACP Journal Club 2000; 132: A11-12.
Glanville J, Lefebvre C, White V, Sheldon T. Searching for systematic reviews in MEDLINE: developing more objective search strategies. 9th Annual Cochrane Colloquium; 2001; Lyon.
Glanville J, Lefebvre C, Wright, K. (editors). ISSG Search Filter Resource 2019a: [updated 2019 Sept 10; cited 2019 Oct 06] https://sites.google.com/a/york.ac.uk/issg-search-filters-resource/home.
Glanville J, Foxlee R, Wisniewski S, Noel-Storr A, Edwards M, Dooley G. Translating the Cochrane EMBASE RCT filter from the Ovid interface to Embase.com: a case study. Health Information and Libraries Journal 2019b; 36: 264-277.
Glanville J, Dooley G, Wisniewski S, Foxlee R, Noel-Storr A. Development of a search filter to identify reports of controlled clinical trials within CINAHL Plus. Health Information and Libraries Journal 2019c; 36: 73-90.
Glanville JM, Lefebvre C, Miles JN, Camosso-Stefinovic J. How to identify randomized controlled trials in MEDLINE: ten years on. Journal of the Medical Library Association 2006; 94: 130-136.
Glanville JM, Duffy S, McCool R, Varley D. Searching ClinicalTrials.gov and the International Clinical Trials Registry Platform to inform systematic reviews: what are the optimal search approaches? Journal of the Medical Library Association 2014; 102: 177-183.
Godin K, Stapleton J, Kirkpatrick SI, Hanning RM, Leatherdale ST. Applying systematic review search methods to the grey literature: a case study examining guidelines for school-based breakfast programs in Canada. Systematic Reviews 2015; 4: 1-10.
Goldacre B, Turner E, on behalf of the OpenTrials team. You can now search FDA approval documents easily at fda.opentrials.net. BMJ 2017; 356: j677.
Goldacre B, DeVito NJ, Heneghan C, Irving F, Bacon S, Fleminger J, Curtis H. Compliance with requirement to report results on the EU Clinical Trials Register: cohort study and web resource. BMJ 2018; 362: k3218.
Gomis M, Gall C, Brahmi FA. Web-based citation management compared to EndNote: options for medical sciences. Medical Reference Services Quarterly 2008; 27: 260-271.
Greenhalgh T, Peacock R. Effectiveness and efficiency of search methods in systematic reviews of complex evidence: audit of primary sources. BMJ 2005; 331: 1064-1065.
Gusenbauer M. Google Scholar to overshadow them all? Comparing the sizes of 12 academic search engines and bibliographic databases. Scientometrics 2019; 118: 177-214.
Haddaway NR, Bayliss HR. Shades of grey: Two forms of grey literature important for reviews in conservation. Biological Conservation 2015; 191: 827-829.
Haddaway NR, Collins AM, Coughlin D, Kirk S. The Role of Google Scholar in Evidence Reviews and Its Applicability to Grey Literature Searching. PloS One 2015; 10: e0138237.
Hannink G, Gooszen HG, Rovers MM. Comparison of registered and published primary outcomes in randomized clinical trials of surgical interventions. Annals of Surgery 2013; 257: 818-823.
HarmoniSR Working Group, for the Cochrane Information Specialists' Executive. HarmoniSR: final project report 2015. http://community.cochrane.org/sites/default/files/uploads/inline-files/HarmoniSR%20-%20final%20report%20-Sept%202015_22-09-2015.pdf.
Hartling L, Featherstone R, Nuspl M, Shave K, Dryden DM, Vandermeer B. Grey literature in systematic reviews: a cross-sectional study of the contribution of non-English reports, unpublished studies and dissertations to the results of meta-analyses in child-relevant reviews. BMC Medical Research Methodology 2017; 17: 64.
Hartung DM, Zarin DA, Guise JM, McDonagh M, Paynter R, Helfand M. Reporting discrepancies between the ClinicalTrials.gov Results Database and peer-reviewed publications. Annals of Internal Medicine 2014; 160: 477-483.
Harzing AW. Publish or Perish 2007. https://harzing.com/resources/publish-or-perish.
Hausner E, Waffenschmidt S, Kaiser T, Simon M. Routine development of objectively derived search strategies. Systematic Reviews 2012; 1: 19.
Hausner E, Guddat C, Hermanns T, Lampert U, Waffenschmidt S. Development of search strategies for systematic reviews: validation showed the noninferiority of the objective approach. Journal of Clinical Epidemiology 2015; 68: 191-199.
Hinde S, Spackman E. Bidirectional citation searching to completion: an exploration of literature searching methods. Pharmacoeconomics 2015; 33: 5-11.
Hocking R. Yale MeSH Analyzer [Product review]. Journal of the Canadian Health Libraries Association 2017; 38: 125-126.
Hodkinson A, Dietz KC, Lefebvre C, Golder S, Jones M, Doshi P, Heneghan C, Jefferson T, Boutron I, Stewart L. The use of clinical study reports to enhance the quality of systematic reviews: a survey of systematic review authors. Systematic Reviews 2018; 7: 117.
Hopewell S, McDonald S, Clarke M, Egger M. Grey literature in meta-analyses of randomized trials of health care interventions. Cochrane Database of Systematic Reviews 2007a; 2: MR000010.
Hopewell S, Clarke M, Lefebvre C, Scherer R. Handsearching versus electronic searching to identify reports of randomized trials. Cochrane Database of Systematic Reviews 2007b; 2: MR000001.
Hopewell S, Clarke MJ, Stewart L, Tierney J. Time to publication for results of clinical trials. Cochrane Database of Systematic Reviews 2007c; 2: MR000011.
Horsley T, Dingwall O, Sampson M. Checking reference lists to find additional studies for systematic reviews. Cochrane Database of Systematic Reviews 2011; 8: MR000026.
Hug SE, Ochsner M, Brandle MP. Citation analysis with microsoft academic. Scientometrics 2017; 111: 371-378.
Hunt DL, McKibbon KA. Locating and Appraising Systematic Reviews. Annals of Internal Medicine 1997; 126: 532-538.
Huser V, Cimino JJ. Linking ClinicalTrials.gov and PubMed to track results of interventional human clinical trials. PloS One 2013a; 8: e68409.
Huser V, Cimino JJ. Evaluating adherence to the International Committee of Medical Journal Editors' policy of mandatory, timely clinical trial registration. Journal of the American Medical Informatics Association 2013b; 20: e169-e174.
Jefferson T, Jones MA, Doshi P, Del Mar CB, Hama R, Thompson MJ, Spencer EA, Onakpoya I, Mahtani KR, Nunan D, Howick J, Heneghan CJ. Neuraminidase inhibitors for preventing and treating influenza in healthy adults and children. Cochrane Database of Systematic Reviews 2014; 4: CD008965.
Jefferson T, Doshi P, Boutron I, Golder S, Heneghan C, Hodkinson A, Jones M, Lefebvre C, Stewart LA. When to include clinical study reports and regulatory documents in systematic reviews. BMJ Evidence-based Medicine 2018; 23: 210-217.
Jiang Y, Lin C, Meng W, Yu C, Cohen AM, Smalheiser NR. Rule-based deduplication of article records from bibliographic databases. Database: the Journal of Biological Databases and Curation 2014; 2014: bat086.
Jones CW, Platts-Mills TF. Quality of registration for clinical trials published in emergency medicine journals. Annals of Emergency Medicine 2012; 60: 458-464.e451.
Jones CW, Handler L, Crowell KE, Keil LG, Weaver MA, Platts-Mills TF. Non-publication of large randomized clinical trials: cross sectional analysis. BMJ 2013; 347: f6104.
Jorgensen L, Gotzsche PC, Jefferson T. Index of the human papillomavirus (HPV) vaccine industry clinical study programmes and non-industry funded studies: a necessary basis to address reporting bias in a systematic review. Systematic Reviews 2018; 7: 8.
Kohl C, McIntosh EJ, Unger S, Haddaway NR, Kecke S, Schiemann J, Wilhelm R. Online tools supporting the conduct and reporting of systematic reviews and systematic maps: a case study on CADIMA and review of existing tools. Environmental Evidence 2018; 7: 8.
Kulkarni AV, Aziz B, Shams I, Busse JW. Comparisons of citations in Web of Science, Scopus, and Google Scholar for articles published in general medical journals. JAMA 2009; 302: 1092-1096.
Kwon Y, Lemieux M, McTavish J, Wathen N. Identifying and removing duplicate records from systematic review searches. Journal of the Medical Library Association 2015; 103: 184-188.
Lanera C, Minto C, Sharma A, Gregori D, Berchialla P, Baldi I. Extending PubMed searches to ClinicalTrials.gov through a machine learning approach for systematic reviews. Journal of Clinical Epidemiology 2018; 103: 22-30.
Lee E, Dobbins M, Decorby K, McRae L, Tirilis D, Husson H. An optimal search filter for retrieving systematic reviews and meta-analyses. BMC Medical Research Methodology 2012; 12: 51.
Lefebvre C, Clarke M. Identifying randomised trials. In: Egger M, Davey Smith G, Altman DG, editors. Systematic Reviews in Health Care: Meta-analysis in Context. 2nd ed. London (UK): BMJ Publication Group; 2001.
Lefebvre C, Eisinga A, McDonald S, Paul N. Enhancing access to reports of randomized trials published world-wide - the contribution of EMBASE records to the Cochrane Central Register of Controlled Trials (CENTRAL) in The Cochrane Library. Emerging Themes in Epidemiology 2008; 5: 13.
Levay P, Raynor M, Tuvey D. The Contributions of MEDLINE, Other Bibliographic Databases and Various Search Techniques to NICE Public Health Guidance. Evidence Based Library and Information Practice 2015; 10: 50-68.
Levay P, Ainsworth N, Kettle R, Morgan A. Identifying evidence for public health guidance: a comparison of citation searching with Web of Science and Google Scholar. Research Synthesis Methods 2016; 7: 34-45.
Lexchin J, Herder M, Doshi P. Canada finally opens up data on new drugs and devices. BMJ 2019; 365: l1825.
Liberati A, Altman DG, Tetzlaff J, Mulrow C, Gotzsche PC, Ioannidis JPA, Clarke M, Devereaux PJ, Kleijnen J, Moher D. The PRISMA Statement for Reporting Systematic Reviews and Meta-Analyses of Studies That Evaluate Health Care Interventions: Explanation and Elaboration. PLoS Medicine 2009; 6: e1000100.
Linder SK, Kamath GR, Pratt GF, Saraykar SS, Volk RJ. Citation searches are more sensitive than keyword searches to identify studies using specific measurement instruments. Journal of Clinical Epidemiology 2015; 68: 412-417.
Littlewood A, Bridges C, for the Cochrane Information Specialist Support Team. Cochrane Information Specialists' Handbook. The Cochrane Collaboration; 2017. http://training.cochrane.org/resource/cochrane-information-specialists-handbook.
Lorenzetti DL, Ghali WA. Reference management software for systematic reviews and meta-analyses: an exploration of usage and usability. BMC Medical Research Methodology 2013; 13: 141.
Lunny C, McKenzie JE, McDonald S. Retrieval of overviews of systematic reviews in MEDLINE was improved by the development of an objectively derived and validated search strategy. Journal of Clinical Epidemiology 2015; 74: 107-118.
Mahood Q, Van Eerd D, Irvin E. Searching for grey literature for systematic reviews: challenges and benefits. Research Synthesis Methods 2014; 5: 221-234.
Manriquez JJ. Searching the LILACS database could improve systematic reviews in dermatology. Archives of Dermatology 2009; 145: 947-948.
Marshall IJ, Noel-Storr A, Kuiper J, Thomas J, Wallace BC. Machine learning for identifying Randomized Controlled Trials: an evaluation and practitioner's guide. Research Synthesis Methods 2018; 9: 602-614.
McDonald S. Improving access to the international coverage of reports of controlled trials in electronic databases: a search of the Australasian Medical Index. Health Information and Libraries Journal 2002; 19: 14-20.
Montori VM, Wilczynski NL, Morgan D, Haynes RB, for the Hedges Team. Optimal search strategies for retrieving systematic reviews from Medline: analytical survey. BMJ 2005; 330: 68.
Moseley AM, Sherrington C, Elkins MR, Herbert RD, Maher CG. Indexing of randomised controlled trials of physiotherapy interventions: a comparison of AMED, CENTRAL, CINAHL, EMBASE, Hooked on Evidence, PEDro, PsycINFO and PubMed. Physiotherapy 2009; 95: 151-156.
Nasser M, Al Hajeri A. A comparison of handsearching versus EMBASE searching of the Archives of Iranian Medicine to identify reports of randomized controlled trials. Archives of Iranian Medicine 2006; 9: 192-195.
Noel-Storr A, Featherstone R, Glanville J, Wisniewski S, Dooley G, Thomas J, Foxlee R. Evaluating Cochrane's centralised search and screening processes: a retrospective analysis of the Cochrane Central Register of Controlled Trial's (CENTRAL) coverage. 26th Cochrane Colloquium; 2019; Santiago, Chile. https://colloquium2019.cochrane.org/abstracts/evaluating-cochranes-centralised-search-and-screening-processes-retrospective-analysis.
O'Mara-Eves A, Brunton G, McDaid D, Kavanagh J, Oliver S, Thomas J. Techniques for identifying cross-disciplinary and ‘hard-to-detect’ evidence for systematic review. Research Synthesis Methods 2014; 5: 50-59.
O'Mara-Eves A, Thomas J, McNaught J, Miwa M, Ananiadou S. Using text mining for study identification in systematic reviews: a systematic review of current approaches. Systematic Reviews 2015; 4: 5.
Ogilvie D, Hamilton V, Egan M, Petticrew M. Systematic reviews of health effects of social interventions: 1. Finding the evidence: how far should you go? Journal of Epidemiology and Community Health 2005; 59: 804-808.
Oxman A, Chalmers I, Clarke M, Enkin M, Schulz K, Starr M, Dickersin K, Herxheimer A, Silagy C (editors). Cochrane Handbook for Systematic Reviews of Interventions. Cochrane Collaboration; 1994.
Paez A. Gray literature: An important resource in systematic reviews. Journal of Evidence-Based Medicine 2017; 10: 233-240.
Page MJ, Shamseer L, Tricco AC. Registration of systematic reviews in PROSPERO: 30,000 records and counting. Systematic Reviews 2018; 7: 32.
Papaioannou D, Sutton A, Carroll C, Booth A, Wong R. Literature searching for social science systematic reviews: consideration of a range of search techniques. Health Information and Libraries Journal 2010; 27: 114-122.
Paynter R, Banez L, Berliner E, Erinoff E, Lege-Matsuura J, Potter S, Uhl S. EPC Methods: An exploration of the use of text-mining software in systematic reviews [Internet]. Rockville (MD): Agency for Healthcare Research and Quality (US); 2016. https://www.ncbi.nlm.nih.gov/books/NBK362044/.
Petticrew M, Song F, Wilson P, Wright K. Quality-assessed reviews of health care interventions and the database of abstracts of reviews of effectiveness (DARE). International Journal of Technology Assessment in Health Care 1999; 15: 671-678.
Pitkin RM, Branagan MA, Burmeister LF. Accuracy of data in abstracts of published research articles. JAMA 1999; 281: 1110-1111.
Rathbone J, Hoffmann T, Glasziou P. Faster title and abstract screening? Evaluating Abstrackr, a semi-automated online screening program for systematic reviewers. Systematic Reviews 2015; 4: 80.
Rathbone J, Carter M, Hoffmann T, Glasziou P. A comparison of the performance of seven key bibliographic databases in identifying all relevant systematic reviews of interventions for hypertension. Systematic Reviews 2016; 5: 27.
Richards D. Handsearching still a valuable element of the systematic review. Evidence-Based Dentistry 2008; 9: 85.
Rogers M, Bethel A, Talens-Bou J, Briscoe S. Chasing references: a comparison of Scopus, Web of Science and Google Scholar for forward citation searching. CILIP Health Libraries Group Conference; 2016; Scarborough.
Rogers M, Bethel A, Abbott R. Locating qualitative studies in dementia on MEDLINE, EMBASE, CINAHL, and PsycINFO: A comparison of search strategies. Research Synthesis Methods 2017; 9: 579-586.
Royle P, Waugh N. Should systematic reviews include searches for published errata? Health Information and Libraries Journal 2004; 21: 14-20.
Royle PL, Bain L, Waugh NR. Sources of evidence for systematic reviews of interventions in diabetes. Diabetic Medicine 2005; 22: 1386-1393.
Salami K, Alkayed K. Publication bias in pediatric hematology and oncology: analysis of abstracts presented at the annual meeting of the American Society of Pediatric Hematology and Oncology. Pediatric Hematology and Oncology 2013; 30: 165-169.
Saleh AA, Ratajeski MA, Bertolet M. Grey Literature Searching for Health Sciences Systematic Reviews: A Prospective Study of Time Spent and Resources Utilized. Evidence Based Library and Information Practice 2014; 9: 28-50.
Sampson M, McGowan J, Cogo E, Horsley T. Managing database overlap in systematic reviews using Batch Citation Matcher: case studies using Scopus. Journal of the Medical Library Association 2006; 94: 461-463.
Sampson M, de Bruijn B, Urquhart C, Shojania K. Complementary approaches to searching MEDLINE may be sufficient for updating existing systematic reviews. Journal of Clinical Epidemiology 2016; 78: 108-115.
Scherer RW, Meerpohl JJ, Pfeifer N, Schmucker C, Schwarzer G, von Elm E. Full publication of results initially presented in abstracts. Cochrane Database of Systematic Reviews 2018; 11: MR000005.
Schmucker CM, Blumle A, Schell LK, Schwarzer G, Oeller P, Cabrera L, von Elm E, Briel M, Meerpohl JJ, on behalf of the OPEN consortium. Systematic review finds that study data not published in full text articles have unclear impact on meta-analyses results in medical research. PloS One 2017; 12: e0176210.
Schoonbaert D. SPIRS, WinSPIRS, and OVID: a comparison of three MEDLINE-on-CD-ROM interfaces [see comment]. Bulletin of the Medical Library Association 1996; 84: 63-70.
Scopus. Scopus: content coverage guide 2017. https://www.elsevier.com/__data/assets/pdf_file/0007/69451/0597-Scopus-Content-Coverage-Guide-US-LETTER-v4-HI-singles-no-ticks.pdf.
Shokraneh F, Adams CE. Study-based registers of randomized controlled trials: Starting a systematic review with data extraction or meta-analysis. BioImpacts 2017; 7: 209-217.
Sinha A, Shen Z, Song Y, Ma H, Elde D, Hsu B-J, Wang K. An overview of Microsoft Academic Service (MAS) and applications. Proceedings of the 24th International Conference on World Wide Web (WWW '15 Companion); 2015; Florence. http://dx.doi.org/10.1145/2740908.2742839.
Slobogean GP, Verma A, Giustini D, Slobogean BL, Mulpuri K. MEDLINE, EMBASE, and Cochrane index most primary studies but not abstracts included in orthopedic meta-analyses. Journal of Clinical Epidemiology 2009; 62: 1261-1267.
Stansfield C, Brunton G, Rees R. Search wide, dig deep: literature searching for qualitative research. An analysis of the publication formats and information sources used for four systematic reviews in public health. Research Synthesis Methods 2014; 5: 142-151.
Stansfield C, Dickson K, Bangpan M. Exploring issues in the conduct of website searching and other online sources for systematic reviews: how can we be systematic? Systematic Reviews 2016; 5: 191.
Stansfield C, O'Mara-Eves A, Thomas J. Text mining for search term development in systematic reviewing: A discussion of some methods and challenges. Research Synthesis Methods 2017; 8: 355-365.
Stevinson C, Lawlor DA. Searching multiple databases for systematic reviews: added value or diminishing returns? Complementary Therapies in Medicine 2004; 12: 228-232.
Stovold E, Hansen S. Handsearching respiratory conference abstracts: a comparison with abstracts identified by an EMBASE search [Poster]. 19th Cochrane Colloquium; 2011; Madrid. http://2011.colloquium.cochrane.org/scientific-programme/posters-session-1.html.
Subirana M, Sola I, Garcia JM, Gich I, Urrutia G. A nursing qualitative systematic review required MEDLINE and CINAHL for study identification. Journal of Clinical Epidemiology 2005; 58: 20-25.
van Driel ML, De Sutter A, De Maeseneer J, Christiaens T. Searching for unpublished trials in Cochrane reviews may not be worth the effort. Journal of Clinical Epidemiology 2009; 62: 838-844.
Vickers AJ, Smith C. Incorporating data from dissertations in systematic reviews. International Journal of Technology Assessment in Health Care 2000; 16: 711-713.
Waffenschmidt S, Hausner E, Kaiser T. An evaluation of searching the German CCMed database for the production of systematic reviews. Health Information and Libraries Journal 2010; 27: 262-267.
Wallace BC, Noel-Storr A, Marshall IJ, Cohen AM, Smalheiser NR, Thomas J. Identifying reports of randomized controlled trials (RCTs) via a hybrid machine learning and crowdsourcing approach. Journal of the American Medical Informatics Association 2017; 24: 1165-1168.
Wanner A, Baumann N. Design and implementation of a tool for conversion of search strategies between PubMed and Ovid MEDLINE. Research Synthesis Methods 2018; 10: 154-160.
Watson RJ, Richardson PH. Identifying randomized controlled trials of cognitive therapy for depression: comparing the efficiency of Embase, Medline and PsycINFO bibliographic databases. British Journal of Medical Psychology 1999a; 72: 535-542.
Watson RJ, Richardson PH. Accessing the literature on outcome studies in group psychotherapy: the sensitivity and precision of Medline and PsycINFO bibliographic database searching. British Journal of Medical Psychology 1999b; 72: 127-134.
Web of Science. Web of Science platform: Web of Science: Summary of Coverage 2019. https://clarivate.libguides.com/webofscienceplatform/coverage.
White VJ, Glanville JM, Lefebvre C, Sheldon TA. A statistical approach to designing search filters to find systematic reviews: objectivity enhances accuracy. Journal of Information Science 2001; 27: 357-370.
Wieland LS, Manheimer E, Sampson M, Barnabas JP, Bouter LM, Cho K, Lee MS, Li X, Liu J, Moher D, Okabe T, Pienaar ED, Shin BC, Tharyan P, Tsutani K, van der Windt DA, Berman BM. Bibliometric and content analysis of the Cochrane Complementary Medicine Field specialized register of controlled trials. Systematic Reviews 2013; 2: 51.
Wilczynski NL, Haynes RB, for the Hedges Team. EMBASE search strategies achieved high sensitivity and specificity for retrieving methodologically sound systematic reviews. Journal of Clinical Epidemiology 2007; 60: 29-33.
Wilczynski NL, Haynes RB. Consistency and accuracy of indexing systematic review articles and meta-analyses in medline. Health Information and Libraries Journal 2009; 26: 203-210.
Wilczynski NL, McKibbon KA, Haynes RB. Sensitive Clinical Queries retrieved relevant systematic reviews as well as primary studies: an analytic survey. Journal of Clinical Epidemiology 2011; 64: 1341-1349.
World Health Organization. Hinari Access to Research for Health programme 2019. http://www.who.int/hinari/en.
Wright K, McDaid C. Reporting of article retractions in bibliographic databases and online journals. Journal of the Medical Library Association 2011; 99: 164-167.
Wright K, Golder S, Rodriguez-Lopez R. Citation searching: a systematic review case study of multiple risk behaviour interventions. BMC Medical Research Methodology 2014; 14: 73.
Wright K, Golder S, Lewis-Light K. What value is the CINAHL database when searching for systematic reviews of qualitative studies? Systematic Reviews 2015; 4: 104.
Wu X-Y, Tang J-L, Mao C, Yuan J-Q, Qin Y, Chung VCH. Systematic reviews and meta-analyses of traditional chinese medicine must search chinese databases to reduce language bias. Evidence-Based Complementary and Alternative Medicine 2013: 812179.
Xia J, Wright J, Adams CE. Five large Chinese biomedical bibliographic databases: accessibility and coverage. Health Information and Libraries Journal 2008; 25: 55-61.
Xue J, Chen W, Chen L, Gaudet L, Moher D, Walker M, Wen SW. Significant discrepancies were found in pooled estimates of searching with Chinese indexes versus searching with English indexes. Journal of Clinical Epidemiology 2016; 70: 246-253.
Younger P, Boddy K. When is a search not a search? A comparison of searching the AMED complementary health database via EBSCOhost, OVID and DIALOG. Health Information and Libraries Journal 2009; 26: 126-135.
Ziai H, Zhang R, Chan A-W, Persaud N. Search for unpublished data by systematic reviewers: an audit. BMJ Open 2017; 7: e017737.
